|
Size: 11289
Comment: Fix suggested by Sharon Robbert
|
Size: 14939
Comment: attachment:HSEL_1.png
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 47: | Line 47: |
| html('<h2>Numerical instability of the classical Gram-Schmidt algorithm</h2>') | pretty_print(html('<h2>Numerical instability of the classical Gram-Schmidt algorithm</h2>')) |
| Line 52: | Line 52: |
| html('precision in bits: ' + str(precision) + '<br>') | pretty_print(html('precision in bits: ' + str(precision) + '<br>')) |
| Line 57: | Line 57: |
| html('Classical Gram-Schmidt:') | pretty_print(html('Classical Gram-Schmidt:')) |
| Line 59: | Line 59: |
| html('Stable Gram-Schmidt:') | pretty_print(html('Stable Gram-Schmidt:')) |
| Line 77: | Line 77: |
| html('<h3>The determinant of a matrix is equal to the determinant of the transpose</h3>') html("$det(%s) = det(%s)=%s$"%(latex(A),latex(A.transpose()),latex(RR(A.determinant())))) |
pretty_print(html('<h3>The determinant of a matrix is equal to the determinant of the transpose</h3>')) pretty_print(html("$\det(%s) = \det(%s)=%s$"%(latex(A),latex(A.transpose()),latex(RR(A.determinant()))))) |
| Line 81: | Line 81: |
| {{attachment:Det_transpose.png}} | {{attachment:Det_transpose_new.png}} |
| Line 94: | Line 94: |
| html("$%s %s=%s$"%tuple(map(latex, [A, v.column().n(4), w.column().n(4)]))) | pretty_print(html("$%s %s=%s$"%tuple(map(latex, [A, v.column().n(4), w.column().n(4)])))) |
| Line 104: | Line 104: |
| html('<h2>The Gerschgorin circle theorem</h2>') | pretty_print(html('<h2>The Gerschgorin circle theorem</h2>')) |
| Line 109: | Line 109: |
| html('$A = ' + latex(matrix(RealField(10),A))+'$') | pretty_print(html('$A = ' + latex(matrix(RealField(10),A))+'$')) |
| Line 173: | Line 173: |
| html('<h3>Singular value decomposition: image of the unit circle and the singular vectors</h3>') html("$A = %s = %s %s %s$"%(latex(my_mat), latex(matrix(rf_low,u.tolist())), latex(matrix(rf_low,2,2,[s[0],0,0,s[1]])), latex(matrix(rf_low,vh.tolist())))) |
pretty_print(html('<h3>Singular value decomposition: image of the unit circle and the singular vectors</h3>')) pretty_print(html("$A = %s = %s %s %s$"%(latex(my_mat), latex(matrix(rf_low,u.tolist())), latex(matrix(rf_low,2,2,[s[0],0,0,s[1]])), latex(matrix(rf_low,vh.tolist()))))) |
| Line 189: | Line 189: |
| html("<h3>Function plot and its discrete Fourier transform</h3>") | pretty_print(html("<h3>Function plot and its discrete Fourier transform</h3>")) |
| Line 225: | Line 225: |
| lista = [(M[j,m],j) for j in range(m+1,D)] | lista = [(abs(M[j,m]),j) for j in range(m+1,D)] |
| Line 257: | Line 257: |
== Solution of an homogeneous system of linear equations == by Pablo Angulo Coefficients are introduced as a matrix in a single text box. The number of equations and unknowns are arbitrary. {{{#!sagecell from sage.misc.html import HtmlFragment def HSLE_as_latex(A, variables): nvars = A.ncols() pretty_print(HtmlFragment( r'$$\left\{\begin{array}{%s}'%('r'*(nvars+1))+ r'\\'.join('%s=&0'%( ' & '.join((r'%s%s\cdot %s'%('+' if c>0 else '',c,v) if c else '') for c,v in zip(row, variables)) if not row.is_zero() else '&'*(nvars-1)+'0' ) for row in A)+ r'\end{array}\right.$$')) @interact def SEL(A='[(0,1,-1,2),(-1,0,2,4), (0,-1,1,-2)]', auto_update=False ): M = A = matrix(eval(A)) neqs = M.nrows() nvars = M.ncols() var_names = ','.join('x%d'%j for j in [1..nvars]) variables = var(var_names) HSLE_as_latex(M, variables) pretty_print(HtmlFragment( 'SEL in matrix form')) show(M) pivot = {} ibound_variables = [] for m,row in enumerate(M): for k in range(m,nvars): lista = [(abs(M[j,k]),j) for j in range(m,neqs)] maxi, c = max(lista) if maxi > 0: ibound_variables.append(k) if M[m,k]==0: M[c,:],M[m,:]=M[m,:],M[c,:] pretty_print( HtmlFragment('We permute rows %d and %d'%(m+1,c+1))) show(M) pivot[m] = k break a=M[m,k] for n in range(m+1,neqs): if M[n,k]!=0: pretty_print( HtmlFragment("We add %s times row %d to row %d"%(-M[n,k]/a, m+1, n+1))) M=M.with_added_multiple_of_row(n,m,-M[n,k]/a) show(M) pretty_print( HtmlFragment('Equivalent system of equations')) HSLE_as_latex(M, variables) SEL_type = 'compatible' null_rows = None for k,row in enumerate(M): if row.is_zero(): pretty_print( HtmlFragment('We remove trivial 0=0 equations')) M = M[:k,:] HSLE_as_latex(M, variables) ifree_variables = [k for k in range(nvars) if k not in ibound_variables] bound_variables = [variables[k] for k in ibound_variables] free_variables = [variables[k] for k in ifree_variables] pretty_print( HtmlFragment('Free variables: %s'% free_variables)) pretty_print( HtmlFragment('Bound variables: %s'% bound_variables)) reduced_eqs = [ sum(c*v for c,v in zip(row, variables))==0 for row in M ] xvector = vector(variables) if len(bound_variables)==1: soldict = solve(reduced_eqs, bound_variables[0], solution_dict=True)[0] else: soldict = solve(reduced_eqs, bound_variables, solution_dict=True)[0] pretty_print( HtmlFragment('Solution in parametric form')) parametric_sol = matrix( xvector.apply_map(lambda s:s.subs(soldict)) ).transpose() show(parametric_sol) pretty_print( HtmlFragment('Generators')) pretty_print( HtmlFragment( r'$$\langle %s\rangle$$'%','.join(latex( parametric_sol.subs(dict((variables[k],1 if j==k else 0) for k in ifree_variables)) ) for j in ifree_variables) )) pretty_print( HtmlFragment('Dimension is %d'%len(free_variables))) }}} {{attachment:HSEL_1.png}} {{attachment:HSEL_2.png}} |
Sage Interactions - Linear Algebra
goto interact main page
Contents
-
Sage Interactions - Linear Algebra
- Numerical instability of the classical Gram-Schmidt algorithm
- Equality of det(A) and det(A.tranpose())
- Linear transformations
- Gerschgorin Circle Theorem
- Singular value decomposition
- Discrete Fourier Transform
- The Gauss-Jordan method for inverting a matrix
- Solution of an homogeneous system of linear equations
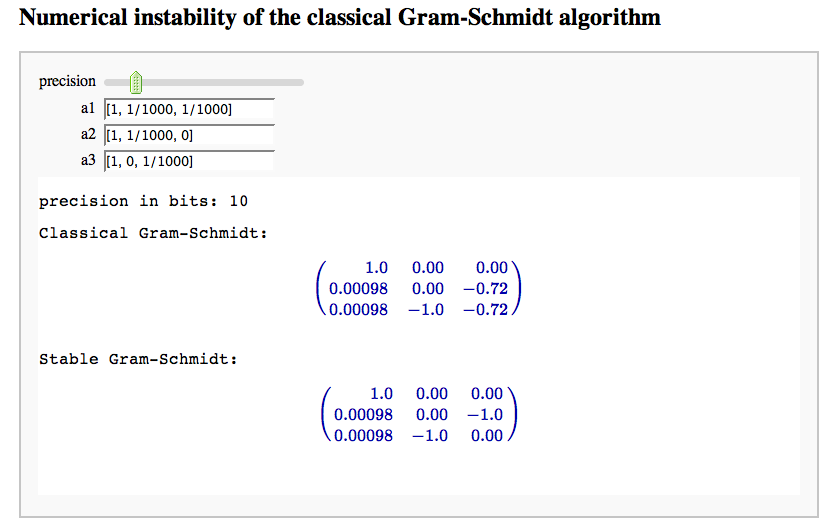
Numerical instability of the classical Gram-Schmidt algorithm
by Marshall Hampton

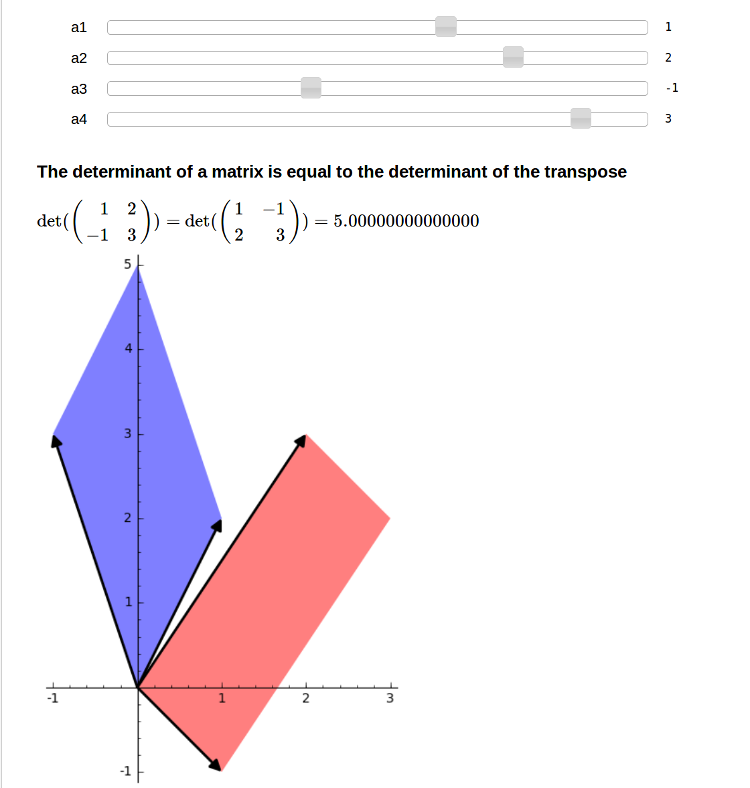
Equality of det(A) and det(A.tranpose())
by Marshall Hampton
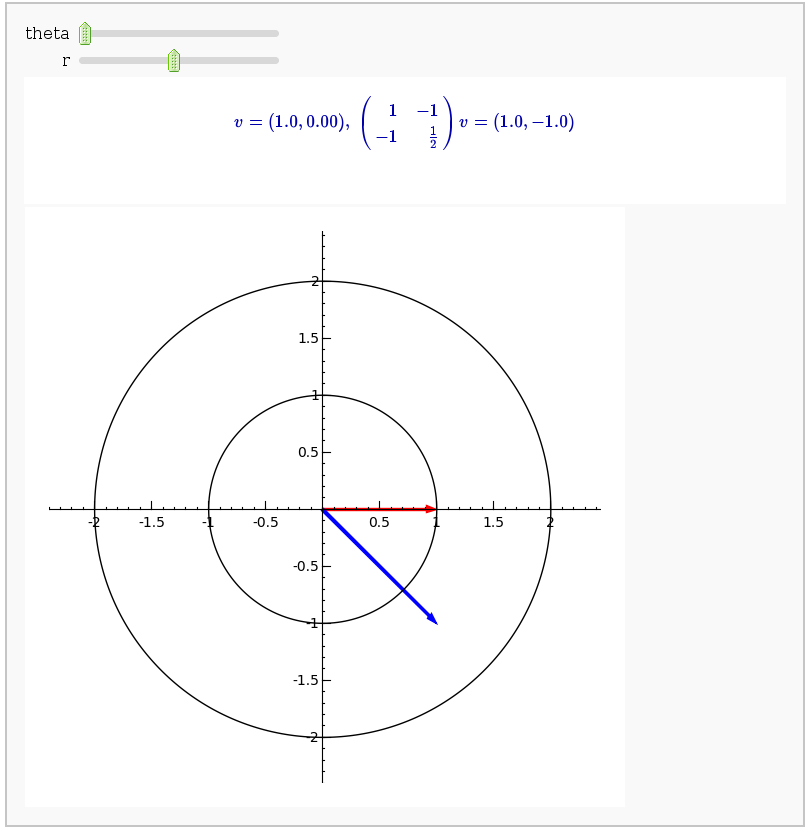
Linear transformations
by Jason Grout
A square matrix defines a linear transformation which rotates and/or scales vectors. In the interact command below, the red vector represents the original vector (v) and the blue vector represents the image w under the linear transformation. You can change the angle and length of v by changing theta and r.
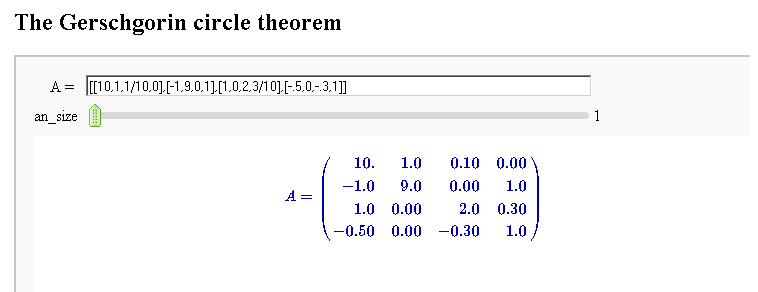
Gerschgorin Circle Theorem
by Marshall Hampton. This animated version requires convert (imagemagick) to be installed, but it can easily be modified to a static version. The animation illustrates the idea behind the stronger version of Gerschgorin's theorem, which says that if the disks around the eigenvalues are disjoint then there is one eigenvalue per disk. The proof is by continuity of the eigenvalues under a homotopy to a diagonal matrix.

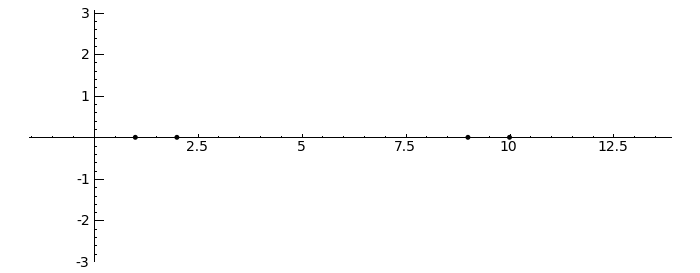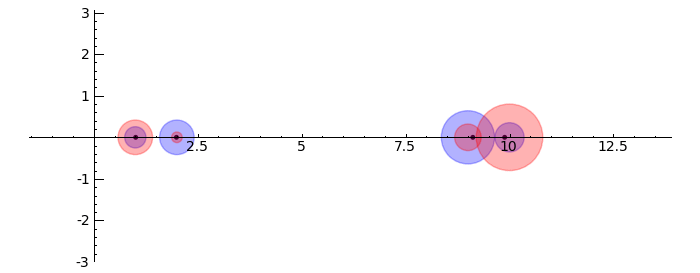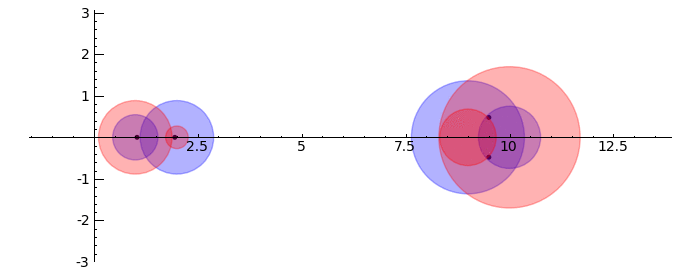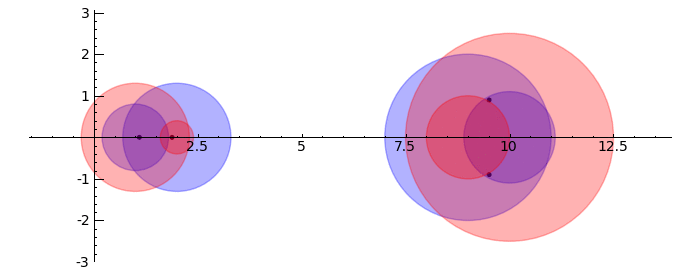
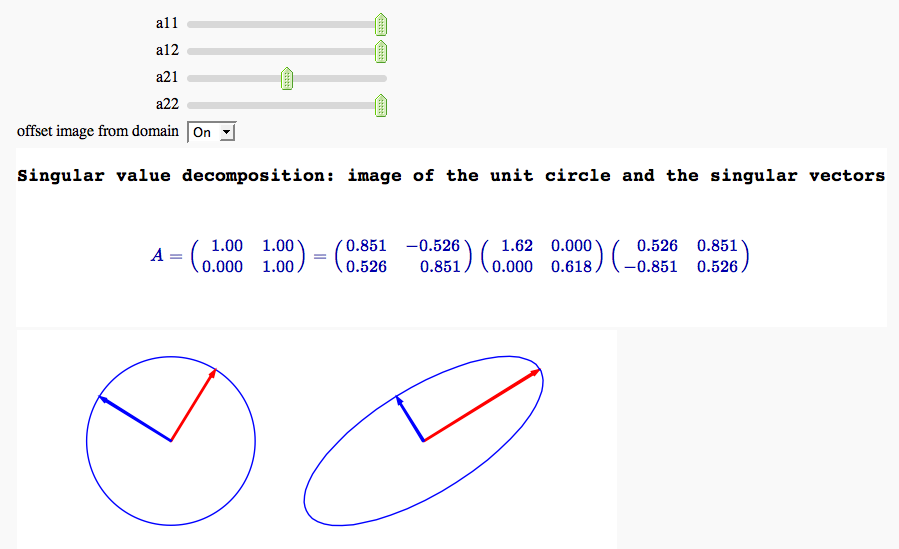
Singular value decomposition
by Marshall Hampton
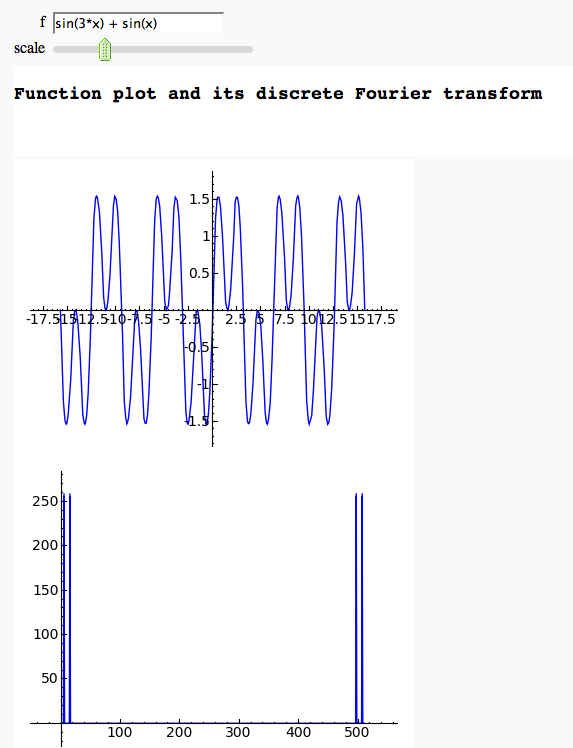
Discrete Fourier Transform
by Marshall Hampton
The Gauss-Jordan method for inverting a matrix
by Hristo Inouzhe
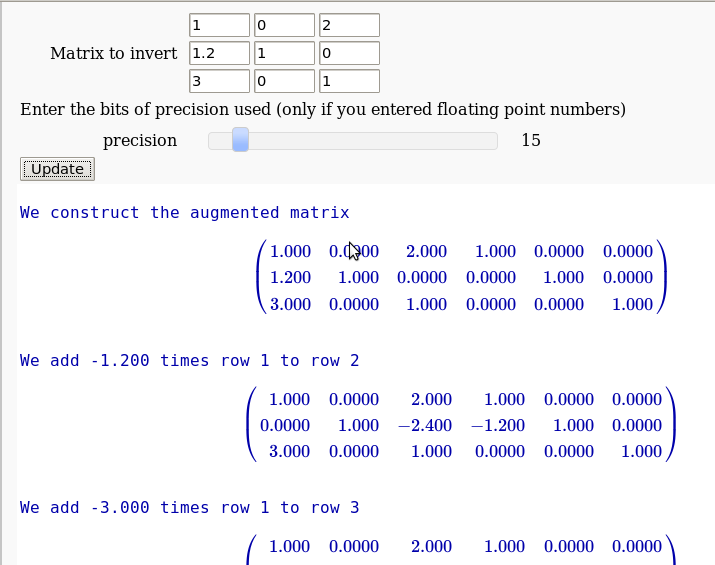
...(goes all the way to invert the matrix)
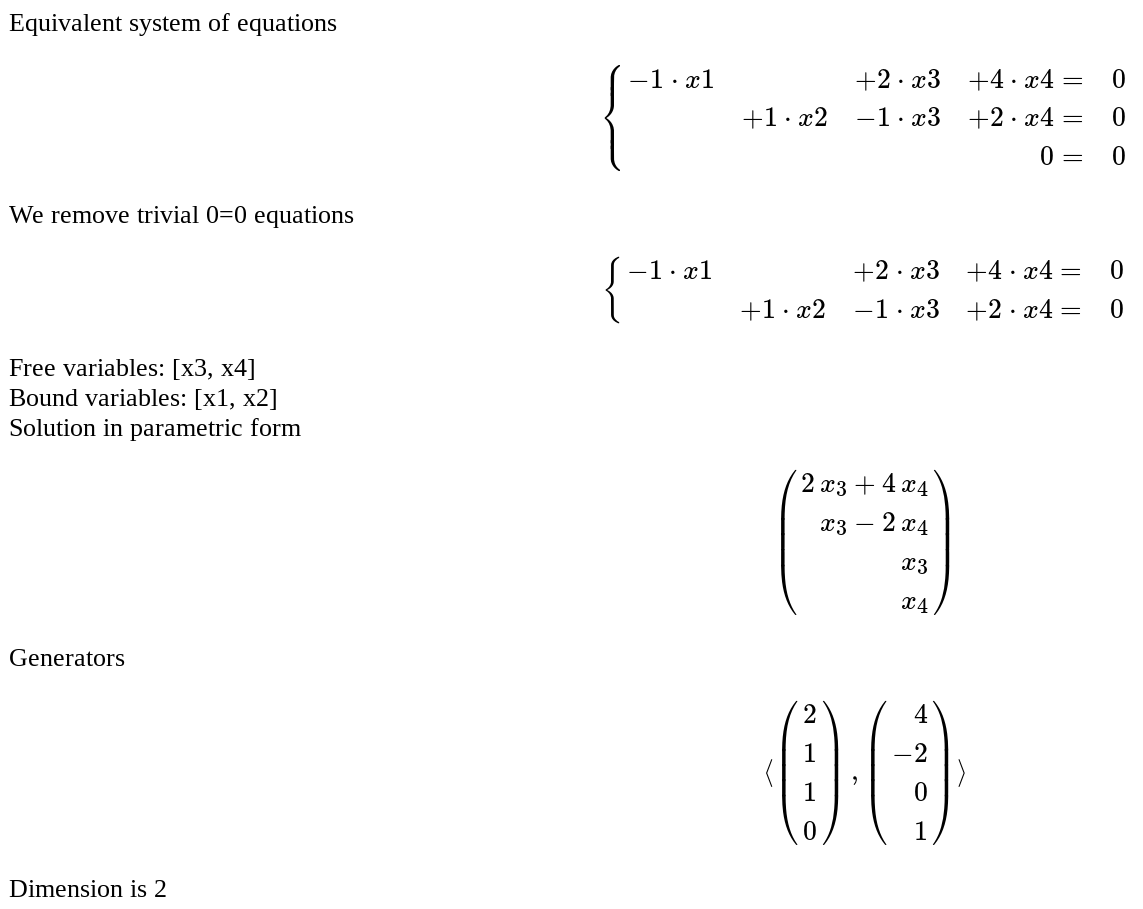
Solution of an homogeneous system of linear equations
by Pablo Angulo
Coefficients are introduced as a matrix in a single text box. The number of equations and unknowns are arbitrary.


