|
Size: 5502
Comment:
|
Size: 19007
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| goto [:interact:interact main page] | goto [[interact|interact main page]] <<TableOfContents>> |
| Line 6: | Line 8: |
| {{{ | {{{#!sagecell |
| Line 28: | Line 30: |
| steps_shown = max(nsteps,show_steps) | steps_shown = min(nsteps,show_steps) |
| Line 36: | Line 38: |
| show(plot(y_exact(x),start,stop,rgbcolor=(1,0,0))+line([[xvals[index],sol[index]] for index in range(len(sol))]),xmin=start,xmax = stop, ymax = sol_max, ymin = sol_min) | show(plot(y_exact(x),start,stop,rgbcolor=(1,0,0))+line([[xvals[index],sol[index]] for index in range(len(sol))]),xmin=start,xmax = stop, ymax = sol_max, ymin = sol_min) |
| Line 43: | Line 45: |
| attachment:eulermethod.png | {{attachment:eulermethod.png}} |
| Line 46: | Line 48: |
| by Mike Hansen (tested and updated by William Stein) {{{ |
by Mike Hansen (tested and updated by William Stein, and later by Dan Drake) {{{#!sagecell |
| Line 49: | Line 51: |
| @interact def _(f = input_box(default=y), g=input_box(default=-x*y+x^3-x), xmin=input_box(default=-1), xmax=input_box(default=1), ymin=input_box(default=-1), ymax=input_box(default=1), start_x=input_box(default=0.5), start_y=input_box(default=0.5), |
from sage.ext.fast_eval import fast_float @interact def _(f = input_box(default=y), g=input_box(default=-x*y+x^3-x), xmin=input_box(default=-1), xmax=input_box(default=1), ymin=input_box(default=-1), ymax=input_box(default=1), start_x=input_box(default=0.5), start_y=input_box(default=0.5), |
| Line 55: | Line 58: |
| old_f = f f = f.function(x,y) old_g = g g = g.function(x,y) |
ff = fast_float(f, 'x', 'y') gg = fast_float(g, 'x', 'y') |
| Line 65: | Line 66: |
| points.append( (xx+step_size*f(xx,yy), yy+step_size*g(xx,yy)) ) | points.append( (xx + step_size * ff(xx,yy), yy + step_size * gg(xx,yy)) ) |
| Line 74: | Line 75: |
| html(r"<h2>$ \frac{dx}{dt} = %s$ $ \frac{dy}{dt} = %s$</h2>"%(latex(old_f),latex(old_g))) print "Step size: %s"%step_size print "Steps: %s"%steps |
html(r"$\displaystyle\frac{dx}{dt} = %s$ $ \displaystyle\frac{dy}{dt} = %s$" % (latex(f),latex(g))) |
| Line 80: | Line 79: |
| attachment:euler.png | {{attachment:euler.png}} |
| Line 84: | Line 83: |
| {{{ | {{{#!sagecell |
| Line 90: | Line 89: |
| def _(fin = input_box(default=y+exp(x/10)-1/3*((x-1/2)^2+y^3)*x-x*y^3), gin=input_box(default=x^3-x+1/100*exp(y*x^2+x*y^2)-0.7*x), xmin=input_box(default=-1), xmax=input_box(default=1.8), ymin=input_box(default=-1.3), ymax=input_box(default=1.5), x_start=(-1,(-2,2)), y_start=(0,(-2,2)), error=(0.5,(0,1)), |
def _(fin = input_box(default=y+exp(x/10)-1/3*((x-1/2)^2+y^3)*x-x*y^3), gin=input_box(default=x^3-x+1/100*exp(y*x^2+x*y^2)-0.7*x), xmin=input_box(default=-1), xmax=input_box(default=1.8), ymin=input_box(default=-1.3), ymax=input_box(default=1.5), x_start=(-1,(-2,2)), y_start=(0,(-2,2)), error=(0.5,(0,1)), |
| Line 107: | Line 106: |
| g(x,y)=gin |
g(x,y)=gin |
| Line 111: | Line 110: |
| Line 113: | Line 112: |
| path = [] | path = [] |
| Line 119: | Line 118: |
| T.function = lambda t, yp: [ff(yp[0],yp[1]), gg(yp[0],yp[1])] | T.function = lambda t, yp: [ff(yp[0],yp[1]), gg(yp[0],yp[1])] |
| Line 129: | Line 128: |
| attachment:ode_runga_kutta.png | {{attachment:ode_runga_kutta.png}} == Linear two-dimensional ODEs == by Marshall Hampton {{{#!sagecell %cython cpdef c_euler_m(double t0, double x10, double x20, double tend, int steps, double a11, double a12, double a21, double a22, double cutoff = 10): cdef double h = (tend-t0)/steps traj = [[x10,x20]] cdef double x1current = x10 cdef double x2current = x20 cdef int i cdef double newx1 cdef double newx2 for i in range(0,steps): newx1 = x1current + h*a11*x1current + h*a12*x2current newx2 = x2current + h*a21*x1current + h*a22*x2current if newx1 > cutoff or newx2 > cutoff or newx1 < -cutoff or newx2 < -cutoff: break traj.append([newx1,newx2]) x1current = newx1 x2current = newx2 return traj }}} {{{#!sagecell @interact def planarsystem(a11 = slider(srange(-10,10,1/10),default = -1), a12 = slider(srange(-10,10,1/10),default = -1), a21 = slider(srange(-10,10,1/10),default = 1), a22 = slider(srange(-10,10,1/10),default = -1), time_tracked = slider(srange(1,100,1.0),default=10)): A = matrix(RDF,[[a11,a12],[a21,a22]]) eigs = A.eigenvalues() html('<center>$x\' = Ax$ dynamics<BR>$A = '+latex(A)+'$, eigenvalues: $%2.2f + %2.2fI, %2.2f + %2.2fI$</center>'%(eigs[0].real(),eigs[0].imag(),eigs[1].real(),eigs[1].imag())) trajs = Graphics() for q in srange(0,2*pi,.15): astart = randint(1,10) ntraj = c_euler_m(0,cos(q),sin(q),time_tracked,300,a11,a12,a21,a22) for i in range(astart,len(ntraj)-1,10): trajs = trajs + arrow(ntraj[i],ntraj[i+1],width=1, arrowsize=2) trajs = trajs + line(ntraj) ntraj = c_euler_m(0,cos(q),sin(q),-time_tracked,300,a11,a12,a21,a22) trajs = trajs + line(ntraj) for i in range(astart,len(ntraj)-1,10): trajs = trajs + arrow(ntraj[i+1],ntraj[i],width=1, arrowsize=2) show(trajs, figsize = [6,6], xmin = -1, xmax = 1, ymin = -1, ymax = 1) }}} {{attachment:linear2x2.png}} == Euler's Method, Improved Euler, and 4th order Runge-Kutta in one variable == by Marshall Hampton. This is a more baroque version of the Euler's method demo above. {{{#!sagecell def ImpEulerMethod(xstart, ystart, xfinish, nsteps, f): sol = [ystart] xvals = [xstart] h = N((xfinish-xstart)/nsteps) for step in range(nsteps): k1 = f(xvals[-1],sol[-1]) k2 = f(xvals[-1] + h,sol[-1] + k1*h) sol.append(sol[-1] + h*(k1+k2)/2) xvals.append(xvals[-1] + h) return zip(xvals,sol) def RK4Method(xstart, ystart, xfinish, nsteps, f): sol = [ystart] xvals = [xstart] h = N((xfinish-xstart)/nsteps) for step in range(nsteps): k1 = f(xvals[-1],sol[-1]) k2 = f(xvals[-1] + h/2,sol[-1] + k1*h/2) k3 = f(xvals[-1] + h/2,sol[-1] + k2*h/2) k4 = f(xvals[-1] + h,sol[-1] + k3*h) sol.append(sol[-1] + h*(k1+2*k2+2*k3+k4)/6) xvals.append(xvals[-1] + h) return zip(xvals,sol) def tab_list(y, headers = None): ''' Converts a list into an html table with borders. ''' s = '<table border = 1>' if headers: for q in headers: s = s + '<th>' + str(q) + '</th>' for x in y: s = s + '<tr>' for q in x: s = s + '<td>' + str(q) + '</td>' s = s + '</tr>' s = s + '</table>' return s var('x, y') @interact def euler_method(q = range_slider(0,10,.1,(0,6),'x range'), y_exact_in = input_box('-cos(x)+2.0', type = str, label = 'Exact solution = '), y_prime_in = input_box('sin(x)', type = str, label = "y' = "), startval = input_box(1.0, label = 'Starting value for y: '), nsteps = slider([2^m for m in range(0,10)], default = 10, label = 'Number of steps: '), show_steps = slider([2^m for m in range(0,10)], default = 8, label = 'Number of steps shown in table: ')): start = q[0] stop = q[1] y_exact = lambda x: eval(y_exact_in) y_prime = lambda x,y: eval(y_prime_in) stepsize = float((stop-start)/nsteps) steps_shown = min(nsteps,show_steps) sol2 = ImpEulerMethod(start, startval, stop, nsteps, y_prime) sol3 = RK4Method(start, startval, stop, nsteps, y_prime) sol = [startval] xvals = [start] for step in range(nsteps): sol.append(sol[-1] + stepsize*y_prime(xvals[-1],sol[-1])) xvals.append(xvals[-1] + stepsize) sol_max = max(sol + [find_maximum_on_interval(y_exact,start,stop)[0]]) sol_min = min(sol + [find_minimum_on_interval(y_exact,start,stop)[0]]) slopes = plot_slope_field(y_prime(x=x,y=y), (start,stop), (sol_min, sol_max)) exact_plot = plot(y_exact(x),start,stop,rgbcolor=(1,0,0)) euler_plot = line([[xvals[index],sol[index]] for index in range(len(sol))]) rk4_plot = line(sol3, rgbcolor = (0,0,.125)) imp_plot = line(sol2, rgbcolor = (1,0,1)) show(slopes +exact_plot + euler_plot + imp_plot+ rk4_plot,xmin=start,xmax = stop, ymax = sol_max, ymin = sol_min) if nsteps < steps_shown: table_range = range(len(sol)) else: table_range = range(0,floor(steps_shown/2)) + range(len(sol)-floor(steps_shown/2),len(sol)) html(tab_list([[i,xvals[i],sol[i],sol2[i][1],sol3[i][1],y_exact(xvals[i])] for i in table_range], headers = ['step','x','<font color="#0000FF">Euler</font>','<font color="#FF00FF">Imp. Euler</font>', '<font color="#0000bb">RK4</font>','<font color="#FF0000">Exact</font>'])) }}} {{attachment:rk4.png}} == Mass/Spring systems == by Jason Grout These two interacts involve some Cython code or other scipy imports, so I've posted a file containing them. You can download the worksheet or copy it online. * https://sage.math.washington.edu/home/pub/42 * https://sage.math.washington.edu/home/pub/43 == Picard iteration example == by Marshall Hampton and David Joyner {{{#!sagecell def picard_iteration(f, a, c, iterations): ''' Computes the N-th Picard iterate for the IVP x' = f(t,x), x(a) = c. EXAMPLES: sage: var('x t s') (x, t, s) sage: a = 0; c = 2 sage: f = lambda t,x: 1-x sage: picard_iteration(f, a, c, 0) 2 sage: picard_iteration(f, a, c, 1) 2 - t sage: picard_iteration(f, a, c, 2) t^2/2 - t + 2 sage: picard_iteration(f, a, c, 3) -t^3/6 + t^2/2 - t + 2 ''' if iterations == 0: return c if iterations == 1: x0 = lambda t: c + integral(f(t=s,x=c), s, a, t) return expand(x0(t)) for i in range(iterations): x_old = lambda s: picard_iteration(f, a, c, iterations-1).subs(t=s) x0 = lambda t: c + integral(f(t=s,x=x_old(s)), s, a, t) return expand(x0(t)) @interact def picarder(n_iterations = slider(0,20,1,default = 2)): var('x,t,s') f = lambda t,x:x html("Exact solution to $x' = x$, $x(0) = 1$ is $x = \exp(t)$<br>") pic = picard_iteration(f,0,1,n_iterations) html('The Picard iteration approximatio after ' + str(n_iterations) + ' iterations is:<br>') html('$'+latex(pic)+'$') exact = plot(exp(t),(t,0,2)) pic_plot = plot(pic,(t,0,2), rgbcolor = (1,0,0)) show(exact + pic_plot) }}} {{attachment:picard.png}} == Euler-Maruyama method and geometric Brownian motion (a common simple model of the stock market) == by Marshall Hampton {{{#!sagecell def EulerMaruyama(xstart, ystart, xfinish, nsteps, f1, f2): sol = [ystart] xvals = [xstart] h = N((xfinish-xstart)/nsteps) for step in range(nsteps): sol.append(sol[-1] + h*f1(sol[-1]) + h^(.5)*f2(sol[-1])*normalvariate(0,1)) xvals.append(xvals[-1] + h) return zip(xvals,sol) out = Graphics() save(out,DATA+'temp') @interact def EulerMaruyamaExample(mu = slider(srange(0,10,.1),default=2.0),sigma = slider(srange(0,10,.1),default=0.5),plots_at_a_time = slider(range(1,100),default=10), number_of_steps = slider(range(1,1000),default=100), clear_plot = checkbox(True), update=selector(['Update'],buttons =True, label='')): html('<center>Example of the Euler-Maruyama method applied to<br>the stochastic differential equation for geometric Brownian motion</center>') html('<center>$S = S_0 + \int_0^t \mu S dt + \int_0^t \sigma S dW$</center>') emplot = list_plot(EulerMaruyama(0,1,1,number_of_steps,lambda x: mu*x,lambda x:sigma*x),plotjoined=True) for i in range(1,plots_at_a_time): emplot = emplot + list_plot(EulerMaruyama(0,1,1,100,lambda x: mu*x,lambda x:sigma*x),plotjoined=True) if clear_plot: out = emplot save(out,DATA+'temp') else: out = load(DATA+'temp') out = out + emplot save(out,DATA+'temp') show(out, figsize = [8,5]) }}} {{attachment:eulermaruyama.png}} == Autonomous equations and stable/unstable fixed points == by Marshall Hampton This needs the Cython functon defined in a seperate cell. Note that it is not a particularly good example of Cython use. {{{#!sagecell %cython cpdef RK4_1d(f, double t_start, double y_start, double t_end, int steps, double y_upper = 10**6, double y_lower = -10**6): ''' Fourth-order scalar Runge-Kutta solver with fixed time steps. f must be a function of t,y, where y is just a scalar variable. ''' cdef double step_size = (t_end - t_start)/steps cdef double t_current = t_start cdef double y_current = y_start cdef list answer_table = [] cdef int j answer_table.append([t_current,y_current]) for j in range(0,steps): k1=f(t_current, y_current) k2=f(t_current+step_size/2, y_current + k1*step_size/2) k3=f(t_current+step_size/2, y_current + k2*step_size/2) k4=f(t_current+step_size, y_current + k3*step_size) t_current += step_size y_current = y_current + (step_size/6)*(k1+2*k2+2*k3+k4) if y_current > y_upper or y_current < y_lower: j = steps answer_table.append([t_current,y_current]) return answer_table }}} {{{#!sagecell from sage.rings.polynomial.real_roots import * var('x') @interact def autonomous_plot(poly=input_box(x*(x-1)*(x-2),label='polynomial'), t_end = slider(1,10,step_size = .1)): var('x') y = polygen(ZZ) ypoly = sage_eval(repr(poly).replace('x','y'),locals=locals()) rr = real_roots(ypoly, max_diameter = 1/100) eps = 0.2 delvals = .04 minr = min([x[0][0] for x in rr]) maxr = max([x[0][1] for x in rr]) svals = [(minr-eps)*t+(1-t)*(maxr+eps) for t in srange(0,1+delvals,delvals)] def polyf(t,xi): return poly(x=xi) paths = [RK4_1d(polyf,0.0,q,t_end,100.0) for q in svals] miny = max(minr-eps,min([min([q1[1] for q1 in q]) for q in paths])) maxy = min(maxr+eps,max([max([q1[1] for q1 in q]) for q in paths])) solpaths = sum([line(q) for q in paths]) fixedpoints = sum([line([[0,(q[0][0]+q[0][1])/2],[t_end,(q[0][0]+q[0][1])/2]], rgbcolor = (1,0,0)) for q in rr]) var('t') html("Autonomous differential equation $x' = p(x)$") show(solpaths+fixedpoints, ymin = miny, ymax = maxy, xmin = 0, xmax = t_end, figsize = [6,4]) }}} {{attachment:Autonomous1.png}} == Heat equation using Fourier series == by Pablo Angulo {{{#!sagecell var('x') x0 = 0 k=1 f = x*exp(-x^2) p = plot(f,0,2*pi, thickness=2) c = 1/pi orden=10 alpha=[(n,c*numerical_integral(f*sin(x*n/2),0,2*pi)[0] ) for n in range(1,orden)] @interact def _(tiempo = (0.1*j for j in (0..10)) ): ft = sum( a*sin(x*n/2)*exp(-k*(n/2)^2*tiempo) for n,a in alpha) pt = plot(ft, 0, 2*pi, color='green', thickness=2) show( p + pt, ymin = -.2) }}} {{attachment:heat_fourier.png}} == Heat equation using finite diferences in cython == by Pablo Angulo {{{#!sagecell %cython #cython code implementing a very simple finite diference scheme import numpy as np def calor_cython(u0,float dx, float k,float t_f,int tsteps): cdef int m cdef float dt cdef float s u=np.array(u0) dt=t_f/tsteps s=k*dt/(dx**2) #we cannot use ^ for exponentiation in cython for m in range(tsteps): u[1:-1]=(1-2*s)*u[1:-1]+s*u[0:-2]+s*u[2:] return u }}} {{{ #interact box wrapping the code above var('x') @interact def _(f=input_box(default=x*exp(-x^2),label='f(x)'), longitud=input_box(default=2*pi), tiempo=input_box(default=0.1), M=input_box(default=100), k=input_box(default=1), tsteps=input_box(default=2000) ): efe=f._fast_float_(x) dx=float(longitud/M) xs=[n*dx for n in range(M+1)] u0=[efe(a) for a in xs] s=k*(tiempo/tsteps) /dx^2 if s>0.5: print 's=%f > 1/2!!! The method is not stable'%s ut=calor_cython(u0,dx,k,tiempo,tsteps) show( line2d(zip(xs, u0)) + line2d(zip(xs, ut), rgbcolor='green') ) }}} {{attachment:heat_findif.png}} == DE with boundary values == The following interact demo looks at the DE+BC y'+y=0, y(0)=a, y(b)=c, and has a slider for b. When b=pi "problems arise":-) {{{#!sagecell var('x') @interact def BCs(b=input_box(1,label='BC at far endpoint'), c = slider(1,5,step_size = .01), a = 1): P1 = text("$y''+y=0,\ y(0) = a,\ y(b) = c$",(4,-1/(c-3.14152))) P2 = plot(a*cos(x)+(b-a*cos(c))*sin(x)/sin(c),x,0,5) (P1+P2).show() }}} [[attachment:de-interact-with-BCs.sws]] |
Sage Interactions - Differential Equations
goto interact main page
Contents
-
Sage Interactions - Differential Equations
- Euler's Method in one variable
- Vector Fields and Euler's Method
- Vector Field with Runga-Kutta-Fehlberg
- Linear two-dimensional ODEs
- Euler's Method, Improved Euler, and 4th order Runge-Kutta in one variable
- Mass/Spring systems
- Picard iteration example
- Euler-Maruyama method and geometric Brownian motion (a common simple model of the stock market)
- Autonomous equations and stable/unstable fixed points
- Heat equation using Fourier series
- Heat equation using finite diferences in cython
- DE with boundary values
Euler's Method in one variable
by Marshall Hampton. This needs some polishing but its usable as is.
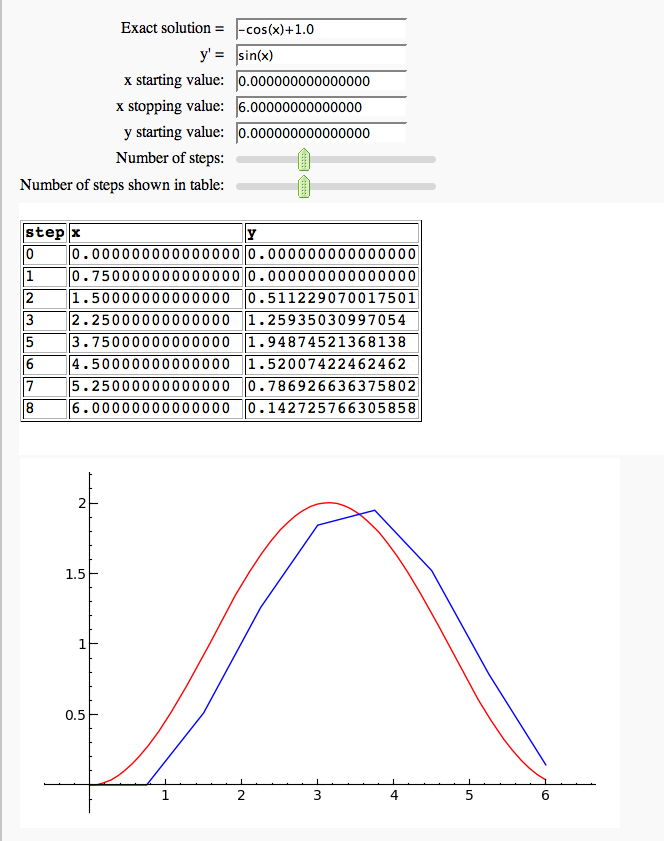
Vector Fields and Euler's Method
by Mike Hansen (tested and updated by William Stein, and later by Dan Drake)
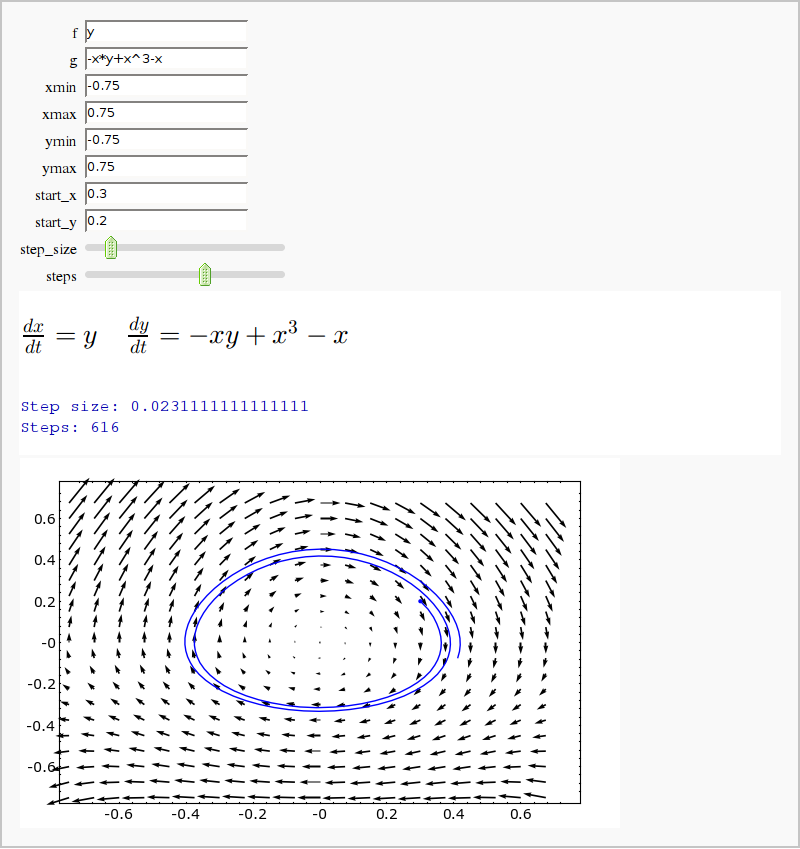
Vector Field with Runga-Kutta-Fehlberg
by Harald Schilly
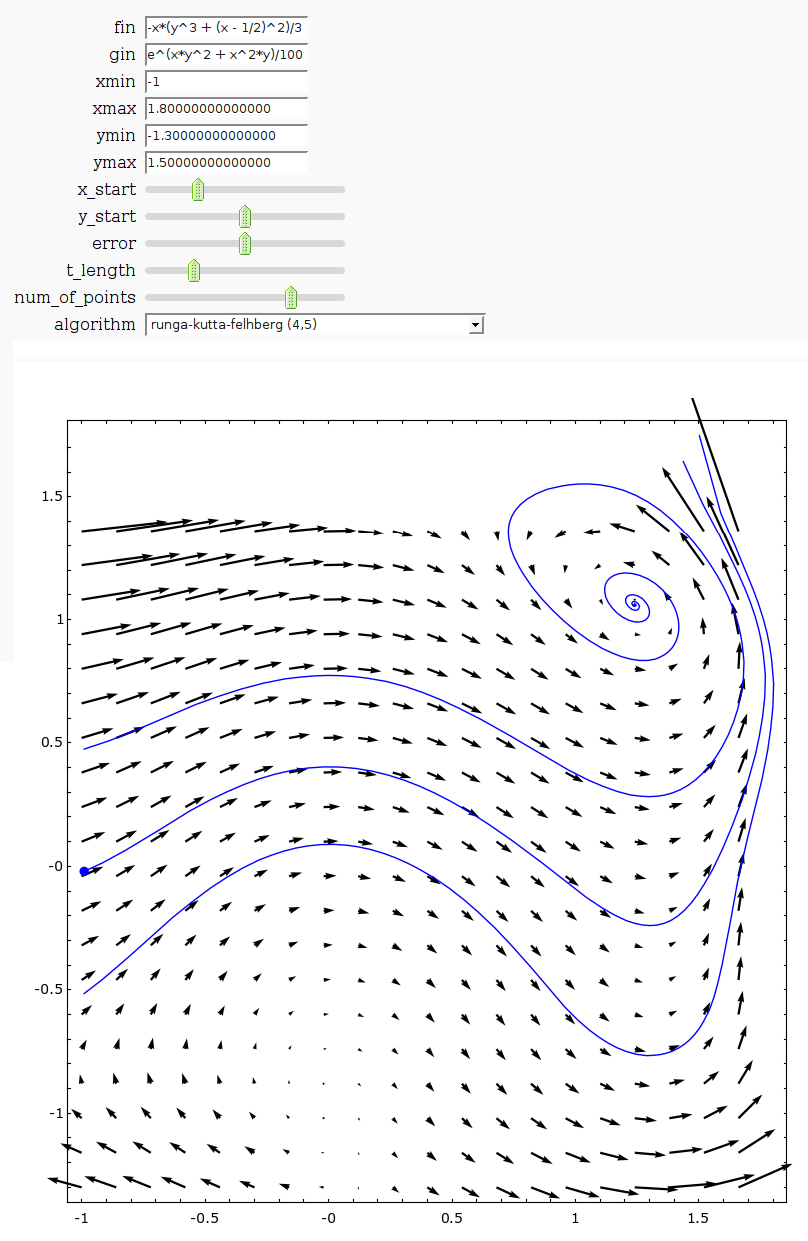
Linear two-dimensional ODEs
by Marshall Hampton
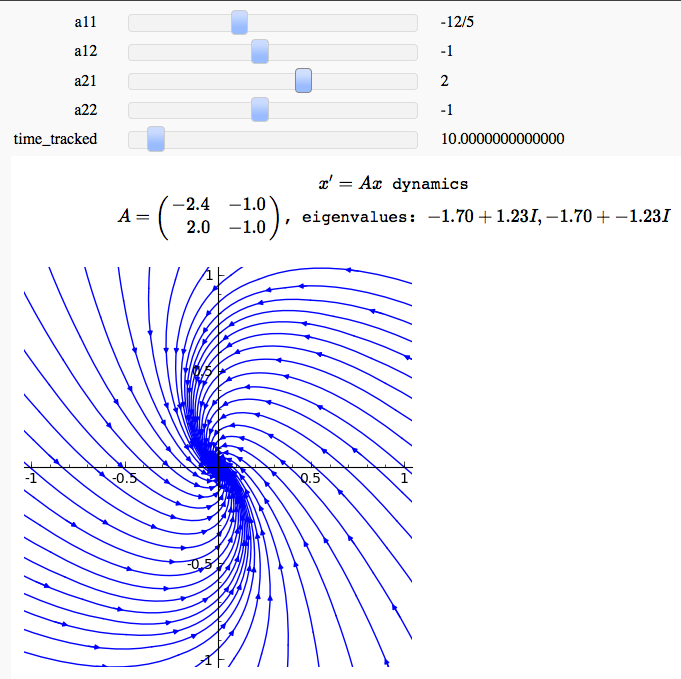
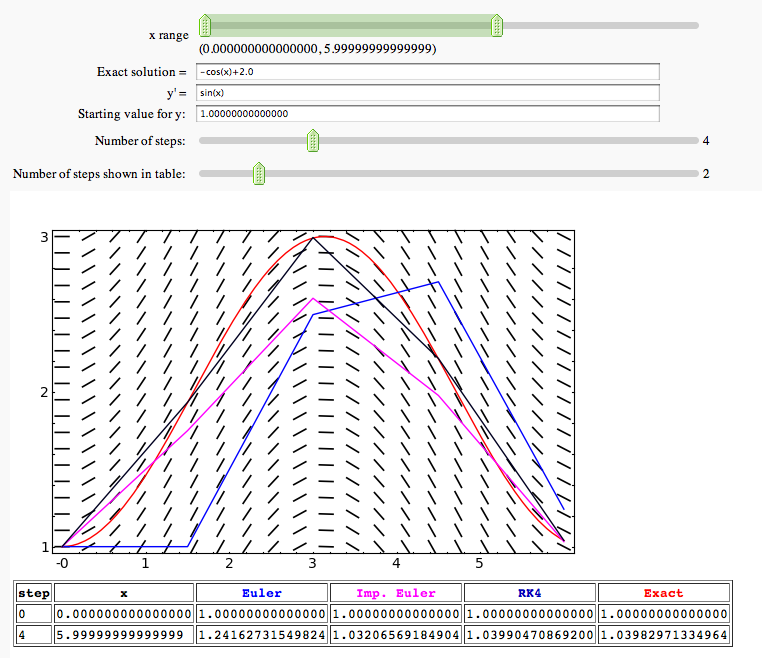
Euler's Method, Improved Euler, and 4th order Runge-Kutta in one variable
by Marshall Hampton. This is a more baroque version of the Euler's method demo above.
Mass/Spring systems
by Jason Grout
These two interacts involve some Cython code or other scipy imports, so I've posted a file containing them. You can download the worksheet or copy it online.
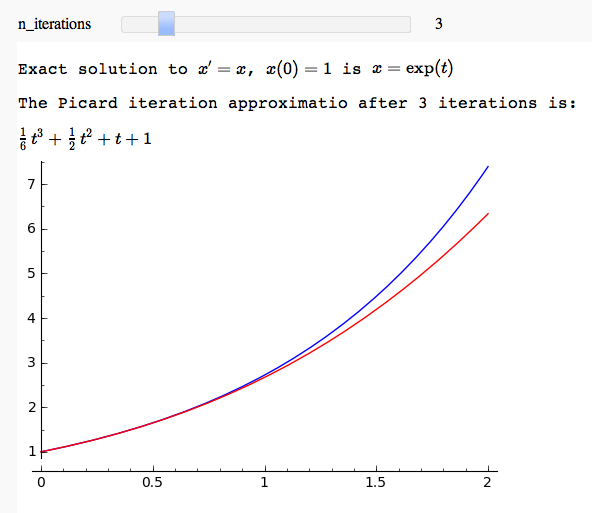
Picard iteration example
by Marshall Hampton and David Joyner
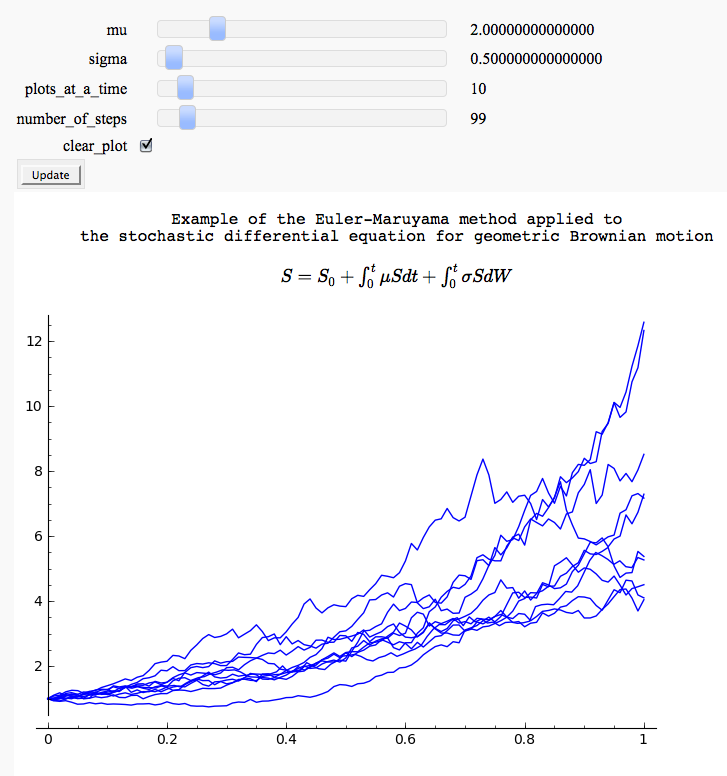
Euler-Maruyama method and geometric Brownian motion (a common simple model of the stock market)
by Marshall Hampton
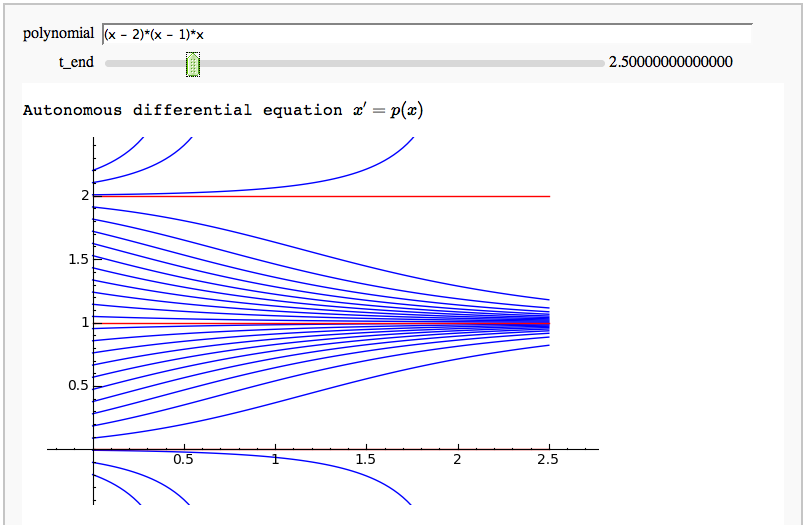
Autonomous equations and stable/unstable fixed points
by Marshall Hampton This needs the Cython functon defined in a seperate cell. Note that it is not a particularly good example of Cython use.
Heat equation using Fourier series
by Pablo Angulo
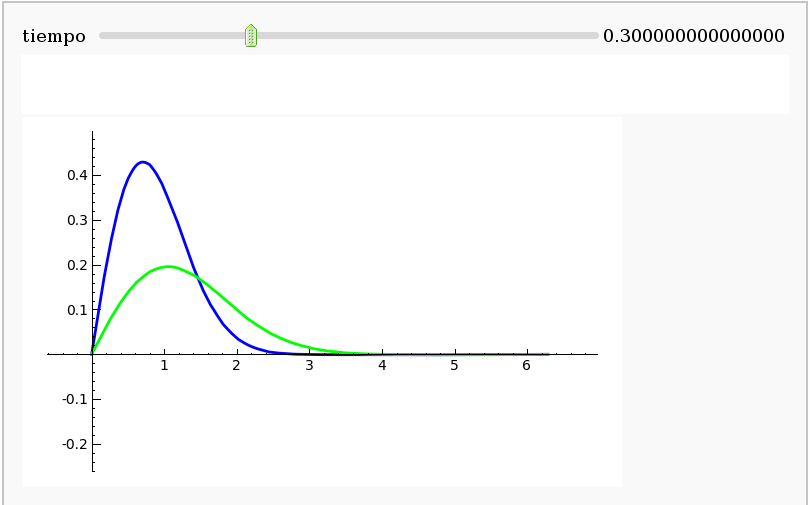
Heat equation using finite diferences in cython
by Pablo Angulo
#interact box wrapping the code above
var('x')
@interact
def _(f=input_box(default=x*exp(-x^2),label='f(x)'), longitud=input_box(default=2*pi),
tiempo=input_box(default=0.1), M=input_box(default=100),
k=input_box(default=1), tsteps=input_box(default=2000) ):
efe=f._fast_float_(x)
dx=float(longitud/M)
xs=[n*dx for n in range(M+1)]
u0=[efe(a) for a in xs]
s=k*(tiempo/tsteps) /dx^2
if s>0.5:
print 's=%f > 1/2!!! The method is not stable'%s
ut=calor_cython(u0,dx,k,tiempo,tsteps)
show( line2d(zip(xs, u0)) + line2d(zip(xs, ut), rgbcolor='green') )
DE with boundary values
The following interact demo looks at the DE+BC y'+y=0, y(0)=a, y(b)=c, and has a slider for b. When b=pi "problems arise":-)
