|
Size: 13991
Comment:
|
Size: 63005
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| goto [:interact:interact main page] [[TableOfContents]] |
goto [[interact|interact main page]] <<TableOfContents>> |
| Line 8: | Line 8: |
| Line 10: | Line 9: |
| {{{ | {{{#!sagecell |
| Line 12: | Line 12: |
| try: | try: |
| Line 19: | Line 19: |
| c = (a+b)/two | c = (a+b)/two |
| Line 27: | Line 27: |
| raise ValueError, "f must have a sign change in the interval (%s,%s)"%(a,b) | raise ValueError("f must have a sign change in the interval (%s,%s)"%(a,b)) |
| Line 29: | Line 29: |
html("<h1>Double Precision Root Finding Using Bisection</h1>") @interact def _(f = cos(x) - x, a = float(0), b = float(1), eps=(-3,(-16..-1))): |
pretty_print(html("<h1>Double Precision Root Finding Using Bisection</h1>")) @interact def _(f = cos(x) - x, a = float(0), b = float(1), eps=(-3,(-16, -1))): |
| Line 34: | Line 33: |
| print "eps = %s"%float(eps) | print("eps = %s" % float(eps)) |
| Line 36: | Line 35: |
| time c, intervals = bisect_method(f, a, b, eps) | c, intervals = bisect_method(f, a, b, eps) |
| Line 38: | Line 37: |
| print "f must have opposite sign at the endpoints of the interval" | print("f must have opposite sign at the endpoints of the interval") |
| Line 41: | Line 40: |
| print "root =", c print "f(c) = %r"%f(c) print "iterations =", len(intervals) |
print("root =", c) print("f(c) = %r" % f(x=c)) print("iterations =", len(intervals)) |
| Line 51: | Line 50: |
| attachment:bisect.png |
{{attachment:bisect.png}} |
| Line 55: | Line 53: |
| Note that there is a more complicated Newton's method below. | |
| Line 57: | Line 56: |
| {{{ | https://cloud.sagemath.com/projects/19575ea0-317e-402b-be57-368d04c113db/files/pub/2801-2901/2824-Double%20Precision%20Root%20Finding%20Using%20Newton's%20Method.sagews {{{#!sagecell |
| Line 61: | Line 63: |
| try: | try: |
| Line 67: | Line 69: |
| for i in xrange(maxiter): | for i in range(maxiter): |
| Line 71: | Line 73: |
| iterates.append(c) return c, iterates html("<h1>Double Precision Root Finding Using Newton's Method</h1>") @interact def _(f = x^2 - 2, c = float(0.5), eps=(-3,(-16..-1)), interval=float(0.5)): |
iterates.append(c) return c, iterates var('x') pretty_print(html("<h1>Double Precision Root Finding Using Newton's Method</h1>")) @interact def _(f = x^2 - 2, c = float(0.5), eps=(-3,(-16, -1)), interval=float(0.5)): |
| Line 81: | Line 81: |
| print "eps = %s"%float(eps) time z, iterates = newton_method(f, c, eps) print "root =", z print "f(c) = %r"%f(z) |
print("eps = %s"%float(eps)) z, iterates = newton_method(f, c, eps) print("root = {}".format(z)) print("f(c) = %r" % f(x=z)) |
| Line 86: | Line 86: |
| print "iterations =", n html(iterates) P = plot(f, z-interval, z+interval, rgbcolor='blue') |
print("iterations = {}".format(n)) pretty_print(html(iterates)) P = plot(f, (x,z-interval, z+interval), rgbcolor='blue') |
| Line 94: | Line 94: |
attachment:newton.png |
{{attachment:newton.png}} |
| Line 99: | Line 98: |
| {{{ @interact def _(q1=(-1,(-3,3)), q2=(-2,(-3,3)), cmap=['autumn', 'bone', 'cool', 'copper', 'gray', 'hot', 'hsv', |
https://cloud.sagemath.com/projects/19575ea0-317e-402b-be57-368d04c113db/files/pub/2801-2901/2823.sagews {{{#!sagecell @interact def _(q1=(-1,(-3,3)), q2=(-2,(-3,3)), cmap=['autumn', 'bone', 'cool', 'copper', 'gray', 'hot', 'hsv', |
| Line 106: | Line 108: |
| C = contour_plot(f, (-2,2), (-2,2), plot_points=30, contours=15, cmap=cmap) | C = contour_plot(f, (x,-2,2), (y,-2,2), plot_points=30, contours=15, cmap=cmap) |
| Line 108: | Line 110: |
| show(plot3d(f, (x,-2,2), (y,-2,2)), figsize=5, viewer='tachyon') }}} attachment:mountains.png |
show(plot3d(f, (x,-2,2), (y,-2,2)), figsize=5, viewer='tachyon') }}} {{attachment:mountains.png}} |
| Line 114: | Line 116: |
| {{{ html('<h2>Tangent line grapher</h2>') |
{{{#!sagecell pretty_print(html('<h2>Tangent line grapher</h2>')) |
| Line 122: | Line 125: |
| tanf = f(x0i) + df(x0i)*(x-x0i) | tanf = f(x=x0i) + df(x=x0i)*(x-x0i) |
| Line 124: | Line 127: |
| print 'Tangent line is y = ' + tanf._repr_() | print('Tangent line is y = ' + tanf._repr_()) |
| Line 126: | Line 129: |
| fmax = f.find_maximum_on_interval(prange[0], prange[1])[0] fmin = f.find_minimum_on_interval(prange[0], prange[1])[0] |
fmax = f.find_local_maximum(prange[0], prange[1])[0] fmin = f.find_local_minimum(prange[0], prange[1])[0] |
| Line 130: | Line 133: |
| attachment:tangents.png | {{attachment:tangents.png}} == Numerical integrals with the midpoint rule == by Marshall Hampton {{{#!sagecell var('x') @interact def midpoint(n = slider(1,100,1,4), f = input_box(default = "x^2", type = str), start = input_box(default = "0", type = str), end = input_box(default = "1", type = str)): a = N(start) b = N(end) func = sage_eval(f, locals={'x':x}) dx = (b-a)/n midxs = [q*dx+dx/2 + a for q in range(n)] midys = [func(x=x_val) for x_val in midxs] rects = Graphics() for q in range(n): xm = midxs[q] ym = midys[q] rects = rects + line([[xm-dx/2,0],[xm-dx/2,ym],[xm+dx/2,ym],[xm+dx/2,0]], rgbcolor = (1,0,0)) + point((xm,ym), rgbcolor = (1,0,0)) min_y = min(0, find_local_minimum(func,a,b)[0]) max_y = max(0, find_local_maximum(func,a,b)[0]) pretty_print(html('<h3>Numerical integrals with the midpoint rule</h3>')) pretty_print(html(r'$\int_{a}^{b}{f(x) dx} {\approx} \sum_i{f(x_i) \Delta x}$')) print("\n\nSage numerical answer: " + str(integral_numerical(func,a,b,max_points = 200)[0])) print("Midpoint estimated answer: " + str(RDF(dx*sum([midys[q] for q in range(n)])))) show(plot(func,a,b) + rects, xmin = a, xmax = b, ymin = min_y, ymax = max_y) }}} {{attachment:num_int.png}} == Numerical integrals with various rules == by Nick Alexander (based on the work of Marshall Hampton) {{{#!sagecell var('x') @interact def midpoint(f = input_box(default = sin(x^2) + 2, type = SR), interval=range_slider(0, 10, 1, default=(0, 4), label="Interval"), number_of_subdivisions = slider(1, 20, 1, default=4, label="Number of boxes"), endpoint_rule = selector(['Midpoint', 'Left', 'Right', 'Upper', 'Lower'], nrows=1, label="Endpoint rule")): a, b = map(QQ, interval) t = var('t') func = fast_callable(f(x=t), RDF, vars=[t]) dx = ZZ(b-a)/ZZ(number_of_subdivisions) xs = [] ys = [] for q in range(number_of_subdivisions): if endpoint_rule == 'Left': xs.append(q*dx + a) elif endpoint_rule == 'Midpoint': xs.append(q*dx + a + dx/2) elif endpoint_rule == 'Right': xs.append(q*dx + a + dx) elif endpoint_rule == 'Upper': x = find_local_maximum(func, q*dx + a, q*dx + dx + a)[1] xs.append(x) elif endpoint_rule == 'Lower': x = find_local_minimum(func, q*dx + a, q*dx + dx + a)[1] xs.append(x) ys = [ func(x) for x in xs ] rects = Graphics() for q in range(number_of_subdivisions): xm = q*dx + dx/2 + a x = xs[q] y = ys[q] rects += line([[xm-dx/2,0],[xm-dx/2,y],[xm+dx/2,y],[xm+dx/2,0]], rgbcolor = (1,0,0)) rects += point((x, y), rgbcolor = (1,0,0)) min_y = min(0, find_local_minimum(func,a,b)[0]) max_y = max(0, find_local_maximum(func,a,b)[0]) # html('<h3>Numerical integrals with the midpoint rule</h3>') show(plot(func,a,b) + rects, xmin = a, xmax = b, ymin = min_y, ymax = max_y) def cap(x): # print only a few digits of precision if x < 1e-4: return 0 return RealField(20)(x) sum_html = "%s \cdot \\left[ %s \\right]" % (dx, ' + '.join([ "f(%s)" % cap(i) for i in xs ])) num_html = "%s \cdot \\left[ %s \\right]" % (dx, ' + '.join([ str(cap(i)) for i in ys ])) numerical_answer = integral_numerical(func,a,b,max_points = 200)[0] estimated_answer = dx * sum([ ys[q] for q in range(number_of_subdivisions)]) pretty_print(html(r''' <div class="math"> \begin{align*} \int_{a}^{b} {f(x) \, dx} & = %s \\\ \sum_{i=1}^{%s} {f(x_i) \, \Delta x} & = %s \\\ & = %s \\\ & = %s . \end{align*} </div>''' % (numerical_answer, number_of_subdivisions, sum_html, num_html, estimated_answer))) }}} {{attachment:num_int2.png}} == Some polar parametric curves == by Marshall Hampton. This is not very general, but could be modified to show other families of polar curves. {{{#!sagecell @interact def para(n1 = slider(1,5,1,default = 2), n2 = slider(1,5,1,default = 3), a1 = slider(1,10,1/10,6/5), a2 = slider(1,10,1/10,6), b = slider(0,2,1/50,0)): var('t') pretty_print(html('$r=' + latex(b+sin(a1*t)^n1 + cos(a2*t)^n2)+'$')) p = parametric_plot((cos(t)*(b+sin(a1*t)^n1 + cos(a2*t)^n2), sin(t)*(b+sin(a1*t)^n1 + cos(a2*t)^n2)), (t,0, 20*pi), plot_points = 1024, rgbcolor = (0,0,0)) show(p, figsize = [5,5], xmin = -2-b, xmax = 2+b, ymin = -2-b, ymax = 2+b, axes = False) }}} {{attachment:polarcurves1.png}} |
| Line 133: | Line 244: |
| Enter symbolic functions $f$, $g$, and $a$, a range, then click the appropriate button to compute and plot some combination of $f$, $g$, and $a$ along with $f$ and $g$. This is inspired by the Matlab funtool GUI. {{{ |
Enter symbolic functions $f$, $g$, and $a$, a range, then click the appropriate button to compute and plot some combination of $f$, $g$, and $a$ along with $f$ and $g$. This is inspired by the Matlab funtool GUI. {{{#!sagecell |
| Line 139: | Line 248: |
| Line 143: | Line 251: |
| 'f+a', 'f-a', 'f*a', 'f/a', 'f^a', 'f(x+a)', 'f(x*a)', 'f+g', 'f-g', 'f*g', 'f/g', 'f(g)'], |
'f+a', 'f-a', 'f*a', 'f/a', 'f^a', 'f(x+a)', 'f(x*a)', 'f+g', 'f-g', 'f*g', 'f/g', 'f(g)'], |
| Line 147: | Line 255: |
| Line 150: | Line 257: |
| except TypeError, msg: print msg[-200:] print "Unable to make sense of f,g, or a as symbolic expressions." return |
except TypeError as msg: print(msg[-200:]) print("Unable to make sense of f,g, or a as symbolic expressions.") return |
| Line 217: | Line 324: |
| html('<center><font color="red">$f = %s$</font></center>'%latex(f)) html('<center><font color="green">$g = %s$</font></center>'%latex(g)) html('<center><font color="blue"><b>$h = %s = %s$</b></font></center>'%(lbl, latex(h))) |
pretty_print(html('<center><font color="red">$f = %s$</font></center>'%latex(f))) pretty_print(html('<center><font color="green">$g = %s$</font></center>'%latex(g))) pretty_print(html('<center><font color="blue"><b>$h = %s = %s$</b></font></center>'%(lbl, latex(h)))) |
| Line 232: | Line 337: |
attachment:funtool.png |
{{attachment:funtool.png}} |
| Line 236: | Line 340: |
| Line 239: | Line 342: |
| This allows user to display the Newton-Raphson procedure one step at a time. It uses the heuristic that, if any of the values of the controls change, then the procedure should be re-started, else it should be continued. {{{ |
This allows user to display the Newton-Raphson procedure one step at a time. It uses the heuristic that, if any of the values of the controls change, then the procedure should be re-started, else it should be continued. {{{#!sagecell |
| Line 247: | Line 348: |
| State = Data = None # globals to allow incremental additions to graphics | State = Data = None # globals to allow incremental changes in interaction data |
| Line 255: | Line 356: |
| step = ['Next','Reset'] ): | step = ['Next','Prev', 'Reset'] ): |
| Line 257: | Line 358: |
| prange = [xmin,xmax] | |
| Line 260: | Line 360: |
| X = [RR(x0)] # restart the plot df = diff(f) Fplot = plot(f, prange[0], prange[1]) Data = [X, df, Fplot] |
Data = [ 1 ] # reset the plot |
| Line 265: | Line 362: |
X, df, Fplot = Data i = len(X) - 1 # compute and append the next x value xi = X[i] fi = RR(f(xi)) fpi = RR(df(xi)) xip1 = xi - fi/fpi X.append(xip1) msg = xip1s = None # now check x value for reasonableness |
elif step == 'Next': N, = Data Data = [ N+1 ] elif step == 'Prev': N, = Data if N > 1: Data = [ N-1 ] N, = Data df = diff(f) theplot = plot( f, xmin, xmax ) theplot += text( '\n$x_0$', (x0,0), rgbcolor=(1,0,0), vertical_alignment="bottom" if f(x=x0) < 0 else "top" ) theplot += points( [(x0,0)], rgbcolor=(1,0,0) ) Trace = [] def Err( msg, Trace=Trace ): Trace.append( '<font color="red"><b>Error: %s!!</b></font>' % (msg,) ) def Disp( s, color="blue", Trace=Trace ): Trace.append( """<font color="%s">$ %s $</font>""" % (color,s,) ) Disp( """f(x) = %s""" % (latex(f),) ) Disp( """f'(x) = %s""" % (latex(df),) ) stop = False |
| Line 276: | Line 388: |
| if abs(xip1) > 10E6*(xmax-xmin): is_inf = True show_calcs = True msg = 'Derivative is 0!' xip1s = latex(xip1.sign()*infinity) X.pop() elif not ((xmin - 0.5*(xmax-xmin)) <= xip1 <= (xmax + 0.5*(xmax-xmin))): show_calcs = True msg = 'x value out of range; probable divergence!' if xip1s is None: xip1s = '%.4g' % (xip1,) def Disp( s, color="blue" ): if show_calcs: html( """<font color="%s">$ %s $</font>""" % (color,s,) ) Disp( """f(x) = %s""" % (latex(f),) + """~~~~f'(x) = %s""" % (latex(df),) ) Disp( """i = %d""" % (i,) + """~~~~x_{%d} = %.4g""" % (i,xi) + """~~~~f(x_{%d}) = %.4g""" % (i,fi) + """~~~~f'(x_{%d}) = %.4g""" % (i,fpi) ) if msg: html( """<font color="red"><b>%s</b></font>""" % (msg,) ) c = "red" else: c = "blue" Disp( r"""x_{%d} = %.4g - ({%.4g})/({%.4g}) = %s""" % (i+1,xi,fi,fpi,xip1s), color=c ) Fplot += line( [(xi,0),(xi,fi)], linestyle=':', rgbcolor=(1,0,0) ) # vert dotted line Fplot += points( [(xi,0),(xi,fi)], rgbcolor=(1,0,0) ) labi = text( '\nx%d\n' % (i,), (xi,0), rgbcolor=(1,0,0), vertical_alignment="bottom" if fi < 0 else "top" ) if is_inf: xl = xi - 0.05*(xmax-xmin) xr = xi + 0.05*(xmax-xmin) yl = yr = fi else: xl = min(xi,xip1) - 0.02*(xmax-xmin) xr = max(xi,xip1) + 0.02*(xmax-xmin) yl = -(xip1-xl)*fpi yr = (xr-xip1)*fpi Fplot += points( [(xip1,0)], rgbcolor=(0,0,1) ) # new x value labi += text( '\nx%d\n' % (i+1,), (xip1,0), rgbcolor=(1,0,0), vertical_alignment="bottom" if fi < 0 else "top" ) Fplot += line( [(xl,yl),(xr,yr)], rgbcolor=(1,0,0) ) # tangent show( Fplot+labi, xmin = prange[0], xmax = prange[1] ) Data = [X, df, Fplot] }}} attachment:newtraph.png == Coordinate Transformations == |
xi = x0 for i in range(N): fi = RR(f(x=xi)) fpi = RR(df(x=xi)) theplot += points( [(xi,fi)], rgbcolor=(1,0,0) ) theplot += line( [(xi,0),(xi,fi)], linestyle=':', rgbcolor=(1,0,0) ) # vert dotted line Disp( """i = %d""" % (i,) ) Disp( """~~~~x_{%d} = %.4g""" % (i,xi) ) Disp( """~~~~f(x_{%d}) = %.4g""" % (i,fi) ) Disp( """~~~~f'(x_{%d}) = %.4g""" % (i,fpi) ) if fpi == 0.0: Err( 'Derivative is 0 at iteration %d' % (i+1,) ) is_inf = True show_calcs = True else: xip1 = xi - fi/fpi Disp( r"""~~~~x_{%d} = %.4g - ({%.4g})/({%.4g}) = %.4g""" % (i+1,xi,fi,fpi,xip1) ) if abs(xip1) > 10*(xmax-xmin): Err( 'Derivative is too close to 0!' ) is_inf = True show_calcs = True elif not ((xmin - 0.5*(xmax-xmin)) <= xip1 <= (xmax + 0.5*(xmax-xmin))): Err( 'x value out of range; probable divergence!' ) stop = True show_calcs = True if is_inf: xl = xi - 0.05*(xmax-xmin) xr = xi + 0.05*(xmax-xmin) yl = yr = fi else: xl = min(xi,xip1) - 0.01*(xmax-xmin) xr = max(xi,xip1) + 0.01*(xmax-xmin) yl = -(xip1-xl)*fpi yr = (xr-xip1)*fpi theplot += text( '\n$x_{%d}$' % (i+1,), (xip1,0), rgbcolor=(1,0,0), vertical_alignment="bottom" if f(x=xip1) < 0 else "top" ) theplot += points( [(xip1,0)], rgbcolor=(1,0,0) ) theplot += line( [(xl,yl),(xr,yr)], rgbcolor=(1,0,0) ) # tangent if stop or is_inf: break epsa = 100.0*abs((xip1-xi)/xip1) nsf = 2 - log(2.0*epsa)/log(10.0) Disp( r"""~~~~~~~~\epsilon_a = \left|(%.4g - %.4g)/%.4g\right|\times100\%% = %.4g \%%""" % (xip1,xi,xip1,epsa) ) Disp( r"""~~~~~~~~num.~sig.~fig. \approx %.2g""" % (nsf,) ) xi = xip1 show( theplot, xmin=xmin, xmax=xmax ) if show_calcs: for t in Trace: pretty_print(html( t )) }}} {{attachment:newtraph.png}} == Coordinate Transformations (FIXME in Jupyter) == |
| Line 333: | Line 449: |
| {{{ | {{{#!sagecell |
| Line 335: | Line 452: |
| # polar coordinates #(x,y)=(u*cos(v),u*sin(v)); (u_range,v_range)=([0..6],[0..2*pi,step=pi/12]) # weird example (x,y)=(u^2-v^2,u*v+cos(u*v)); (u_range,v_range)=([-5..5],[-5..5]) thickness=4 square_length=.05 |
|
| Line 336: | Line 462: |
| @interact def trans(x=input_box(u^2-v^2, label="x=",type=SR), \ y=input_box(u*v, label="y=",type=SR), \ t_val=slider(0,10,0.2,6, label="Length of curves"), \ u_percent=slider(0,1,0.05,label="<font color='red'>u</font>", default=.7), v_percent=slider(0,1,0.05,label="<font color='blue'>v</font>", default=.7), u_range=input_box(range(-5,5,1), label="u lines"), v_range=input_box(range(-5,5,1), label="v lines")): thickness=4 |
from functools import partial @interact def trans(x=input_box(x, label="x",type=SR), y=input_box(y, label="y",type=SR), u_percent=slider(0,1,0.05,label="<font color='red'>u</font>", default=.7), v_percent=slider(0,1,0.05,label="<font color='blue'>v</font>", default=.7), t_val=slider(0,10,0.2,6, label="Length"), u_range=input_box(u_range, label="u lines"), v_range=input_box(v_range, label="v lines")): x(u,v)=x y(u,v)=y |
| Line 349: | Line 478: |
g1=sum([parametric_plot((SR(u.subs(u=i))._fast_float_('v'),v.subs(u=i)._fast_float_('v')), t_min,t_max, rgbcolor=(1,0,0)) for i in u_range]) g2=sum([parametric_plot((u.subs(v=i)._fast_float_('u'),SR(v.subs(v=i))._fast_float_('u')), t_min,t_max, rgbcolor=(0,0,1)) for i in v_range]) vline_straight=parametric_plot((SR(u.subs(v=v_val))._fast_float_('u'),SR(v.subs(v=v_val))._fast_float_('u')), t_min,t_max, rgbcolor=(0,0,1), linestyle='-',thickness=thickness) uline_straight=parametric_plot((SR(u.subs(u=u_val))._fast_float_('v'),SR(v.subs(u=u_val))._fast_float_('v')), t_min,t_max,rgbcolor=(1,0,0), linestyle='-',thickness=thickness) (g1+g2+vline_straight+uline_straight).save("uv_coord.png",aspect_ratio=1, figsize=[5,5], axes_labels=['$u$','$v$']) g3=sum([parametric_plot((x.subs(u=i)._fast_float_('v'),y.subs(u=i)._fast_float_('v')), t_min,t_max, rgbcolor=(1,0,0)) for i in u_range]) g4=sum([parametric_plot((x.subs(v=i)._fast_float_('u'),y.subs(v=i)._fast_float_('u')), t_min,t_max, rgbcolor=(0,0,1)) for i in v_range]) vline=parametric_plot((SR(x.subs(v=v_val))._fast_float_('u'),SR(y.subs(v=v_val))._fast_float_('u')), t_min,t_max, rgbcolor=(0,0,1), linestyle='-',thickness=thickness) uline=parametric_plot((SR(x.subs(u=u_val))._fast_float_('v'),SR(y.subs(u=u_val))._fast_float_('v')), t_min,t_max,rgbcolor=(1,0,0), linestyle='-',thickness=thickness) (g3+g4+vline+uline).save("xy_coord.png", aspect_ratio=1, figsize=[5,5], axes_labels=['$x$','$y$']) print jsmath("x=%s, \: y=%s"%(latex(x), latex(y))) print "<html><table><tr><td><img src='cell://uv_coord.png'/></td><td><img src='cell://xy_coord.png'/></td></tr></table></html>" }}} attachment:coordinate-transform-1.png attachment:coordinate-transform-2.png |
uvplot=sum([parametric_plot((i,v), (v,t_min,t_max), color='red',axes_labels=['u','v'],figsize=[5,5]) for i in u_range]) uvplot+=sum([parametric_plot((u,i), (u,t_min,t_max), color='blue',axes_labels=['u','v']) for i in v_range]) uvplot+=parametric_plot((u,v_val), (u,t_min,t_max), rgbcolor=(0,0,1), linestyle='-',thickness=thickness) uvplot+=parametric_plot((u_val, v), (v,t_min,t_max),rgbcolor=(1,0,0), linestyle='-',thickness=thickness) pt=vector([u_val,v_val]) du=vector([(t_max-t_min)*square_length,0]) dv=vector([0,(t_max-t_min)*square_length]) uvplot+=polygon([pt,pt+dv,pt+du+dv,pt+du],color='purple',alpha=0.7) uvplot+=line([pt,pt+dv,pt+du+dv,pt+du],color='green') T(u,v)=(x,y) xuv = fast_float(x,'u','v') yuv = fast_float(y,'u','v') xvu = fast_float(x,'v','u') yvu = fast_float(y,'v','u') xyplot=sum([parametric_plot((partial(xuv,i),partial(yuv,i)), (v,t_min,t_max), color='red', axes_labels=['x','y'],figsize=[5,5]) for i in u_range]) xyplot+=sum([parametric_plot((partial(xvu,i),partial(yvu,i)), (u,t_min,t_max), color='blue') for i in v_range]) xyplot+=parametric_plot((partial(xuv,u_val),partial(yuv,u_val)),(v,t_min,t_max),color='red', linestyle='-',thickness=thickness) xyplot+=parametric_plot((partial(xvu,v_val),partial(yvu,v_val)), (u,t_min,t_max), color='blue', linestyle='-',thickness=thickness) jacobian=abs(T.diff().det()).simplify_full() t_vals=[0..1,step=t_val*.01] vertices=[(x(*c),y(*c)) for c in [pt+t*dv for t in t_vals]] vertices+=[(x(*c),y(*c)) for c in [pt+dv+t*du for t in t_vals]] vertices+=[(x(*c),y(*c)) for c in [pt+(1-t)*dv+du for t in t_vals]] vertices+=[(x(*c),y(*c)) for c in [pt+(1-t)*du for t in t_vals]] xyplot+=polygon(vertices,color='purple',alpha=0.7) xyplot+=line(vertices,color='green') pretty_print(html("$T(u,v)=%s$"%(latex(T(u,v))))) pretty_print(html("Jacobian: $%s$"%latex(jacobian(u,v)))) pretty_print(html("A very small region in $xy$ plane is approximately %0.4g times the size of the corresponding region in the $uv$ plane"%jacobian(u_val,v_val).n())) pretty_print(table([[uvplot,xyplot]])) }}} {{attachment:coordinate-transform-1.png}} {{attachment:coordinate-transform-2.png}} |
| Line 374: | Line 516: |
| {{{ | {{{#!sagecell |
| Line 379: | Line 522: |
| dot = point((x0,f(x0)),pointsize=80,rgbcolor=(1,0,0)) @interact def _(order=(1..12)): |
dot = point((x0,f(x=x0)),pointsize=80,rgbcolor=(1,0,0)) @interact def _(order=[1..12]): |
| Line 384: | Line 527: |
| html('$f(x)\;=\;%s$'%latex(f)) html('$\hat{f}(x;%s)\;=\;%s+\mathcal{O}(x^{%s})$'%(x0,latex(ft),order+1)) |
pretty_print(html('$f(x)\;=\;%s$'%latex(f))) pretty_print(html('$\hat{f}(x;%s)\;=\;%s+\mathcal{O}(x^{%s})$'%(x0,latex(ft),order+1))) |
| Line 388: | Line 531: |
| attachment:taylor_series_animated.gif | {{attachment:taylor_series_animated.gif}} == Illustration of the precise definition of a limit == by John Perry I'll break tradition and put the image first. Apologies if this is Not A Good Thing. {{attachment:snapshot_epsilon_delta.png}} {{{#!sagecell pretty_print(html("<h2>Limits: <i>ε-δ</i></h2>")) pretty_print(html("This allows you to estimate which values of <i>δ</i> guarantee that <i>f</i> is within <i>ε</i> units of a limit.")) pretty_print(html("<ul><li>Modify the value of <i>f</i> to choose a function.</li>")) pretty_print(html("<li>Modify the value of <i>a</i> to change the <i>x</i>-value where the limit is being estimated.</li>")) pretty_print(html("<li>Modify the value of <i>L</i> to change your guess of the limit.</li>")) pretty_print(html("<li>Modify the values of <i>δ</i> and <i>ε</i> to modify the rectangle.</li></ul>")) pretty_print(html("If the blue curve passes through the pink boxes, your values for <i>δ</i> and/or <i>ε</i> are probably wrong.")) @interact def delta_epsilon(f = input_box(default=(x^2-x)/(x-1), label="$f$"), a=input_box(default=1, label="$a$"), L = input_box(default=1, label="$L$"), delta=input_box(label=r"$\delta$",default=0.1), epsilon=input_box(label=r"$\varepsilon$",default=0.1), xm=input_box(label=r"$x_{min}$",default=-1), xM=input_box(label=r"$x_{max}$",default=4)): f_left_plot = plot(f,xm,a-delta/3,thickness=2) f_right_plot = plot(f,a+delta/3,xM,thickness=2) epsilon_line_1 = line([(xm,L-epsilon),(xM,L-epsilon)], rgbcolor=(0.5,0.5,0.5),linestyle='--') epsilon_line_2 = line([(xm,L+epsilon),(xM,L+epsilon)], rgbcolor=(0.5,0.5,0.5),linestyle='--') ym = min(f_right_plot.ymin(),f_left_plot.ymin()) yM = max(f_right_plot.ymax(),f_left_plot.ymax()) bad_region_1 = polygon([(a-delta,L+epsilon),(a-delta,yM),(a+delta,yM),(a+delta,L+epsilon)], rgbcolor=(1,0.6,0.6)) bad_region_2 = polygon([(a-delta,L-epsilon),(a-delta,ym),(a+delta,ym),(a+delta,L-epsilon)], rgbcolor=(1,0.6,0.6)) aL_point = point((a,L),rgbcolor=(1,0,0),pointsize=20) delta_line_1 = line([(a-delta,ym),(a-delta,yM)],rgbcolor=(0.5,0.5,0.5),linestyle='--') delta_line_2 = line([(a+delta,ym),(a+delta,yM)],rgbcolor=(0.5,0.5,0.5),linestyle='--') (f_left_plot +f_right_plot +epsilon_line_1 +epsilon_line_2 +delta_line_1 +delta_line_2 +aL_point +bad_region_1 +bad_region_2).show(xmin=xm,xmax=xM) }}} == A graphical illustration of sin(x)/x -> 1 as x-> 0 == by Wai Yan Pong {{{#!sagecell x=var('x') @interact def _(x = slider(-7/10,7/10,1/20,1/2)): pretty_print(html('<h3>A graphical illustration of $\lim_{x -> 0} \sin(x)/x =1$</h3>')) pretty_print(html('Below is the unit circle, so the length of the <font color=red>red line</font> is |sin(x)|')) pretty_print(html('and the length of the <font color=blue>blue line</font> is |tan(x)| where x is the length of the arc.')) pretty_print(html('From the picture, we see that |sin(x)| $\le$ |x| $\le$ |tan(x)|.')) pretty_print(html('It follows easily from this that cos(x) $\le$ sin(x)/x $\le$ 1 when x is near 0.')) pretty_print(html('As $\lim_{x ->0} \cos(x) =1$, we conclude that $\lim_{x -> 0} \sin(x)/x =1$.')) if not (x == 0): pretty_print("sin(x)/x = "+str(sin(float(x))/float(x))) elif x == 0: pretty_print("The limit of sin(x)/x as x tends to 0 is 1.") C=circle((0,0),1, rgbcolor='black') mvp = (cos(x),sin(x));tpt = (1, tan(x)) p1 = point(mvp, pointsize=30, rgbcolor='red'); p2 = point((1,0), pointsize=30, rgbcolor='red') line1 = line([(0,0),tpt], rgbcolor='black'); line2 = line([(cos(x),0),mvp], rgbcolor='red') line3 = line([(0,0),(1,0)], rgbcolor='black'); line4 = line([(1,0),tpt], rgbcolor='blue') result = C+p1+p2+line1+line2+line3+line4 result.show(aspect_ratio=1, figsize=[3,3], axes=False) }}} {{attachment:sinelimit.png}} == Quadric Surface Plotter == by Marshall Hampton. This is pretty simple, so I encourage people to spruce it up. In particular, it isn't set up to show all possible types of quadrics. {{{#!sagecell var('x,y,z') quadrics = {'Ellipsoid':x^2+y^2+z^2-1,'Elliptic paraboloid':x^2+y^2-z,'Hyperbolic paraboloid':x^2-y^2-z, '1-Sheeted Hyperboloid':x^2+y^2-z^2-1,'2-Sheeted Hyperboloid':x^2-y^2-z^2-1, 'Cone':x^2+y^2-z^2} @interact def quads(q = selector(list(quadrics)), a = slider(0,5,1/2,default = 1)): f = quadrics[q].subs({x:x*a^(1/2)}) if a==0 or q=='Cone': pretty_print(latex(f), " (degenerate)") else: pretty_print(latex(f)) p = implicit_plot3d(f,(x,-2,2),(y,-2,2),(z,-2,2), plot_points = 75) show(p) }}} {{attachment:quadrics.png}} == The midpoint rule for numerically integrating a function of two variables == by Marshall Hampton {{{#!sagecell from sage.plot.plot3d.platonic import index_face_set def cuboid(v1,v2,**kwds): """ Cuboid defined by corner points v1 and v2. """ ptlist = [] for vi in (v1,v2): for vj in (v1,v2): for vk in (v1,v2): ptlist.append([vi[0],vj[1],vk[2]]) f_incs = [[0, 2, 6, 4], [0, 1, 3, 2], [0, 1, 5, 4], [1, 3, 7, 5], [2, 3, 7, 6], [4, 5, 7, 6]] if 'aspect_ratio' not in kwds: kwds['aspect_ratio'] = [1,1,1] return index_face_set(f_incs,ptlist,enclosed = True, **kwds) var('x,y') R16 = RealField(16) npi = RDF(pi) pretty_print(html("<h1>The midpoint rule for a function of two variables</h1>")) @interact def midpoint2d(func = input_box('y*sin(x)/x+sin(y)',type=str,label='function of x and y'), nx = slider(2,20,1,3,label='x subdivisions'), ny = slider(2,20,1,3,label='y subdivisions'), x_start = slider(-10,10,.1,0), x_end = slider(-10,10,.1,3*npi), y_start= slider(-10,10,.1,0), y_end= slider(-10,10,.1,3*npi)): f = sage_eval('lambda x,y: ' + func) delx = (x_end - x_start)/nx dely = (y_end - y_start)/ny xvals = [RDF(x_start + (i+1.0/2)*delx) for i in range(nx)] yvals = [RDF(y_start + (i+1.0/2)*dely) for i in range(ny)] num_approx = 0 cubs = [] darea = delx*dely for xv in xvals: for yv in yvals: num_approx += f(xv,yv)*darea cubs.append(cuboid([xv-delx/2,yv-dely/2,0],[xv+delx/2,yv+dely/2,f(xv,yv)], opacity = .5, rgbcolor = (1,0,0))) pretty_print(html("$$\int_{"+str(R16(y_start))+"}^{"+str(R16(y_end))+"} "+ "\int_{"+str(R16(x_start))+"}^{"+str(R16(x_end))+"} "+func+"\ dx \ dy$$")) pretty_print(html('<p style="text-align: center;">Numerical approximation: ' + str(num_approx)+'</p>')) p1 = plot3d(f,(x,x_start,x_end),(y,y_start,y_end)) show(p1+sum(cubs)) }}} {{attachment:numint2d.png}} == Gaussian (Legendre) quadrature == by Jason Grout The output shows the points evaluated using Gaussian quadrature (using a weight of 1, so using Legendre polynomials). The vertical bars are shaded to represent the relative weights of the points (darker = more weight). The error in the trapezoid, Simpson, and quadrature methods is both printed out and compared through a bar graph. The "Real" error is the error returned from scipy on the definite integral. {{{#!sagecell import scipy import numpy from scipy.special.orthogonal import p_roots, t_roots, u_roots from scipy.integrate import quad, trapz, simps from sage.ext.fast_eval import fast_float from numpy import linspace show_weight_graph=False # 'Hermite': {'w': e**(-x**2), 'xmin': -numpy.inf, 'xmax': numpy.inf, 'func': h_roots}, # 'Laguerre': {'w': e**(-x), 'xmin': 0, 'xmax': numpy.inf, 'func': l_roots}, methods = {'Legendre': {'w': 1, 'xmin': -1, 'xmax': 1, 'func': p_roots}, 'Chebyshev': {'w': 1/sqrt(1-x**2), 'xmin': -1, 'xmax': 1, 'func': t_roots}, 'Chebyshev2': {'w': sqrt(1-x**2), 'xmin': -1, 'xmax': 1, 'func': u_roots}, 'Trapezoid': {'w': 1, 'xmin': -1, 'xmax': 1, 'func': lambda n: (linspace(-1r,1,n), numpy.array([1.0r]+[2.0r]*(n-2)+[1.0r])*1.0r/n)}, 'Simpson': {'w': 1, 'xmin': -1, 'xmax': 1, 'func': lambda n: (linspace(-1r,1,n), numpy.array([1.0r]+[4.0r,2.0r]*int((n-3.0r)/2.0r)+[4.0r,1.0r])*2.0r/(3.0r*n))}} var("x") def box(center, height, area,**kwds): width2 = 1.0*area/height/2.0 return polygon([(center-width2,0), (center+width2,0),(center+width2,height),(center-width2,height)],**kwds) @interact def weights(n=slider(1,30,1,default=10),f=input_box(default=3*x+cos(10*x),type=SR), show_method=["Legendre", "Chebyshev", "Chebyshev2", "Trapezoid","Simpson"]): ff = fast_float(f,'x') method = methods[show_method] xcoords,w = (method['func'])(int(n)) xmin = method['xmin'] xmax = method['xmax'] plot_min = max(xmin, -10) plot_max = min(xmax, 10) scaled_func = f*method['w'] scaled_ff = fast_float(scaled_func, 'x') coords = zip(xcoords,w) max_weight = max(w) coords_scaled = zip(xcoords,w/max_weight) f_graph = plot(scaled_func,plot_min,plot_max) boxes = sum(box(x,ff(x),w*ff(x),rgbcolor=(0.5,0.5,0.5),alpha=0.3) for x,w in coords) stems = sum(line([(x,0),(x,scaled_ff(x))],rgbcolor=(1-y,1-y,1-y), thickness=2,markersize=6,alpha=y) for x,y in coords_scaled) points = sum([point([(x,0), (x,scaled_ff(x))],rgbcolor='black',pointsize=30) for x,_ in coords]) graph = stems+points+f_graph+boxes if show_weight_graph: graph += line([(x,y) for x,y in coords_scaled], rgbcolor='green',alpha=0.4) show(graph,xmin=plot_min,xmax=plot_max,aspect_ratio="auto") approximation = sum([w*ff(x) for x,w in coords]) integral,integral_error = scipy.integrate.quad(scaled_ff, xmin, xmax) x_val = linspace(min(xcoords), max(xcoords),n) y_val = map(scaled_ff,x_val) trapezoid = integral-trapz(y_val, x_val) simpson = integral-simps(y_val, x_val) pretty_print(html("$$\sum_{i=1}^{i=%s}w_i\left(%s\\right)= %s\\approx %s =\int_{-1}^{1}%s \,dx$$"%(n, latex(f), approximation, integral, latex(scaled_func)))) error_data = [trapezoid, simpson, integral-approximation,integral_error] print("Trapezoid: %s, Simpson: %s, \nMethod: %s, Real: %s" % tuple(error_data)) show(bar_chart(error_data,width=1),ymin=min(error_data), ymax=max(error_data)) }}} {{attachment:quadrature1.png}} {{attachment:quadrature2.png}} == Vector Calculus, 2-D Motion == By Rob Beezer A fast_float() version is available in a [[http://buzzard.ups.edu/sage/motion-2d.sws|worksheet]] {{{#!sagecell # 2-D motion and vector calculus # Copyright 2009, Robert A. Beezer # Creative Commons BY-SA 3.0 US # # 2009/02/15 Built on Sage 3.3.rc0 # 2009/02/17 Improvements from Jason Grout # # variable parameter is t # later at a particular value named t0 # var('t') # # parameter range # start=0 stop=2*pi # # position vector definition # edit here for new example # example is wide ellipse # adjust x, extents in final show() # position=vector( (4*cos(t), sin(t)) ) # # graphic of the motion itself # path = parametric_plot( position.list(), (t, start, stop), color = "black" ) # # derivatives of motion, lengths, unit vectors, etc # velocity = derivative(position, t) acceleration = derivative(velocity, t) speed = velocity.norm() speed_deriv = derivative(speed, t) tangent = (1/speed)*velocity dT = derivative(tangent, t) normal = (1/dT.norm())*dT # # interact section # slider for parameter, 24 settings # checkboxes for various vector displays # computations at one value of parameter, t0 # @interact def _(t0 = slider(float(start), float(stop), float((stop-start)/24), float(start) , label = "Parameter"), pos_check = ("Position", True), vel_check = ("Velocity", False), tan_check = ("Unit Tangent", False), nor_check = ("Unit Normal", False), acc_check = ("Acceleration", False), tancomp_check = ("Tangential Component", False), norcomp_check = ("Normal Component", False) ): # # location of interest # pos_tzero = position(t=t0) # # various scalar quantities at point # speed_component = speed(t=t0) tangent_component = speed_deriv(t=t0) normal_component = sqrt( acceleration(t=t0).norm()^2 - tangent_component^2 ) curvature = normal_component/speed_component^2 # # various vectors, mostly as arrows from the point # pos = arrow((0,0), pos_tzero, rgbcolor=(0,0,0)) tan = arrow(pos_tzero, pos_tzero + tangent(t=t0), rgbcolor=(0,1,0) ) vel = arrow(pos_tzero, pos_tzero + velocity(t=t0), rgbcolor=(0,0.5,0)) nor = arrow(pos_tzero, pos_tzero + normal(t=t0), rgbcolor=(0.5,0,0)) acc = arrow(pos_tzero, pos_tzero + acceleration(t=t0), rgbcolor=(1,0,1)) tancomp = arrow(pos_tzero, pos_tzero + tangent_component*tangent(t=t0), rgbcolor=(1,0,1) ) norcomp = arrow(pos_tzero, pos_tzero + normal_component*normal(t=t0), rgbcolor=(1,0,1)) # # accumulate the graphic based on checkboxes # picture = path if pos_check: picture = picture + pos if vel_check: picture = picture + vel if tan_check: picture = picture+ tan if nor_check: picture = picture + nor if acc_check: picture = picture + acc if tancomp_check: picture = picture + tancomp if norcomp_check: picture = picture + norcomp # # print textual info # print("Position vector defined as r(t)={}".format(position)) print("Speed is {}".format(N(speed(t=t0)))) print("Curvature is {}".format(N(curvature))) # # show accumulated graphical info # adjust x-,y- extents to get best plot # show(picture, xmin=-4,xmax=4, ymin=-1.5,ymax=1.5,aspect_ratio=1) }}} {{attachment:motion2d.png}} == Vector Calculus, 3-D Motion == by Rob Beezer Available as a [[http://buzzard.ups.edu/sage/motion-d.sws|worksheet]] {{{#!sagecell # 3-D motion and vector calculus # Copyright 2009, Robert A. Beezer # Creative Commons BY-SA 3.0 US # # # 2009/02/15 Built on Sage 3.3.rc0 # 2009/02/17 Improvements from Jason Grout # # variable parameter is t # later at a particular value named t0 # # un-comment double hash (##) to get # time-consuming torsion computation # var('t') assume(t, 'real') # # parameter range # start=-4*pi stop=8*pi # # position vector definition # edit here for new example # example is wide ellipse # adjust figsize in final show() to get accurate aspect ratio # a=1/(8*pi) c=(3/2)*a position=vector( (exp(a*t)*cos(t), exp(a*t)*sin(t), exp(c*t)) ) # # graphic of the motion itself # path = parametric_plot3d( position.list(), (t, start, stop), color = "black" ) # # derivatives of motion, lengths, unit vectors, etc # velocity = derivative( position, t) acceleration = derivative(velocity, t) speed = velocity.norm() speed_deriv = derivative(speed, t) tangent = (1/speed)*velocity dT = derivative(tangent, t) normal = (1/dT.norm())*dT binormal = tangent.cross_product(normal) ## dB = derivative(binormal, t) # # interact section # slider for parameter, 24 settings # checkboxes for various vector displays # computations at one value of parameter, t0 # @interact def _(t0 = slider(float(start), float(stop), float((stop-start)/24), float(start) , label = "Parameter"), pos_check = ("Position", True), vel_check = ("Velocity", False), tan_check = ("Unit Tangent", False), nor_check = ("Unit Normal", False), bin_check = ("Unit Binormal", False), acc_check = ("Acceleration", False), tancomp_check = ("Tangential Component", False), norcomp_check = ("Normal Component", False) ): # # location of interest # pos_tzero = position(t=t0) # # various scalar quantities at point # speed_component = speed(t=t0) tangent_component = speed_deriv(t=t0) normal_component = sqrt( acceleration(t=t0).norm()^2 - tangent_component^2 ) curvature = normal_component/speed_component^2 ## torsion = (1/speed_component)*(dB(t0)).dot_product(normal(t0)) # # various vectors, mostly as arrows from the point # pos = arrow3d((0,0,0), pos_tzero, rgbcolor=(0,0,0)) tan = arrow3d(pos_tzero, pos_tzero + tangent(t=t0), rgbcolor=(0,1,0) ) vel = arrow3d(pos_tzero, pos_tzero + velocity(t=t0), rgbcolor=(0,0.5,0)) nor = arrow3d(pos_tzero, pos_tzero + normal(t=t0), rgbcolor=(0.5,0,0)) bin = arrow3d(pos_tzero, pos_tzero + binormal(t=t0), rgbcolor=(0,0,0.5)) acc = arrow3d(pos_tzero, pos_tzero + acceleration(t=t0), rgbcolor=(1,0,1)) tancomp = arrow3d(pos_tzero, pos_tzero + tangent_component*tangent(t=t0), rgbcolor=(1,0,1) ) norcomp = arrow3d(pos_tzero, pos_tzero + normal_component*normal(t=t0), rgbcolor=(1,0,1)) # # accumulate the graphic based on checkboxes # picture = path if pos_check: picture = picture + pos if vel_check: picture = picture + vel if tan_check: picture = picture+ tan if nor_check: picture = picture + nor if bin_check: picture = picture + bin if acc_check: picture = picture + acc if tancomp_check: picture = picture + tancomp if norcomp_check: picture = picture + norcomp # # print textual info # print("Position vector: r(t)=", position) print("Speed is ", N(speed(t=t0))) print("Curvature is ", N(curvature)) ## print("Torsion is ", N(torsion)) print() print("Right-click on graphic to zoom to 400%") print("Drag graphic to rotate") # # show accumulated graphical info # show(picture, aspect_ratio=[1,1,1]) }}} {{attachment:motion3d.png}} == Multivariate Limits by Definition == by John Travis http://sagenb.mc.edu/home/pub/97/ {{{#!sagecell ## An interactive way to demonstrate limits of multivariate functions of the form z = f(x,y) ## ## John Travis ## Mississippi College ## ## Spring 2011 ## # Starting point for radius values before collapsing in as R approaches 0. # Functions ought to be "reasonable" within a circular domain of radius R surrounding # the desired (x_0,y_0). var('x,y,z') Rmin=1/10 Rmax=2 @interact(layout=dict(top=[['f'],['x0'],['y0']], bottom=[['in_3d','curves','R','graphjmol']])) def _(f=input_box((x^2-y^2)/(x^2+y^2),width=30,label='$f(x)$'), R=slider(Rmin,Rmax,1/10,Rmax,label=', $R$'), x0=input_box(0,width=10,label='$x_0$'), y0=input_box(0,width=10,label='$y_0$'), curves=checkbox(default=false,label='Show curves'), in_3d=checkbox(default=false,label='3D'), graphjmol=checkbox(default=true,label='Interactive graph')): if graphjmol: view_method = 'jmol' else: view_method = 'tachyon' # converting f to cylindrical coordinates. g(r,t) = f(x=r*cos(t)+x0,y=r*sin(t)+y0) # Sage graphing transformation used to see the original surface. cylinder = (r*cos(t)+x0,r*sin(t)+y0,z) surface = plot3d(g,(t,0,2*pi),(r,1/100,Rmax),transformation=cylinder,opacity=0.2) # Regraph the surface for smaller and smaller radii controlled by the slider. collapsing_surface = plot3d(g,(t,0,2*pi),(r,1/100,R),transformation=cylinder,rgbcolor=(0,1,0)) G = surface+collapsing_surface pretty_print(html('Enter $(x_0 ,y_0 )$ above and see what happens as $ R \\rightarrow 0 $.')) pretty_print(html('The surface has a limit as $(x,y) \\rightarrow $ ('+str(x0)+','+str(y0)+') if the green region collapses to a point.')) # If checked, add a couple of curves on the surface corresponding to limit as x->x0 for y=x^(3/5), # and as y->y0 for x=y^(3/5). Should make this more robust but perhaps using # these relatively obtuse curves could eliminate problems. if curves: curve_x = parametric_plot3d([x0-t,y0-t^(3/5),f(x=x0-t,y=y0-t^(3/5))],(t,Rmin,Rmax),color='red',thickness=10) curve_y = parametric_plot3d([x0+t^(3/5),y0+t,f(x=x0+t^(3/5),y=y0+t)],(t,Rmin,Rmax),color='red',thickness=10) R2 = Rmin/4 G += arrow((x0-Rmin,y0-Rmin^(3/5),f(x=x0-Rmin,y=y0-Rmin^(3/5))),(x0-R2,y0-R2^(3/5),f(x=x0-R2,y=y0-R2^(3/5))),size=30 ) G += arrow((x0+Rmin^(3/5),y0+Rmin,f(x=x0+Rmin^(3/5),y=y0+Rmin)),(x0+R2^(3/5),y0+R2,f(x=x0+R2^(3/5),y=y0+R2)),size=30 ) limit_x = limit(f(x=x0-t,y=y0-t^(3/5)),t=0) limit_y = limit(f(x=x0+t^(3/5),y=y0+t),t=0) text_x = text3d(limit_x,(x0,y0,limit_x)) text_y = text3d(limit_y,(x0,y0,limit_y)) G += curve_x+curve_y+text_x+text_y pretty_print(html('The red curves represent a couple of trajectories on the surface. If they do not meet, then')) pretty_print(html('there is also no limit. (If computer hangs up, likely the computer can not do these limits.)')) pretty_print(html('\n<center><font color="red">$\lim_{(x,?)\\rightarrow(x_0,y_0)} f(x,y) =%s$</font>'%str(limit_x)+' and <font color="red">$\lim_{(?,y)\\rightarrow(x_0,y_0)} f(x,y) =%s$</font></center>'%str(limit_y))) if in_3d: show(G,stereo="redcyan",viewer=view_method) else: show(G,perspective_depth=true,viewer=view_method) }}} {{attachment:3D_Limit_Defn.png}} {{{#!sagecell ## An interactive way to demonstrate limits of multivariate functions of the form z = f(x,y) ## This one uses contour plots and so will work with functions that have asymptotic behavior. ## ## John Travis ## Mississippi College ## ## Spring 2011 ## # An increasing number of contours for z = f(x,y) are utilized surrounding a desired (x_0,y_0). # A limit can be shown to exist at (x_0,y_0) provided the point stays trapped between adjacent # contour lines as the number of lines increases. If the contours change wildly near the point, # then a limit does not exist. # Looking for two different paths to approach (x_0,y_0) that utilize a different selection of colors # will help locate paths to use that exhibit the absence of a limit. var('x,y,z,u') @interact(layout=dict(top=[['f'],['x0'],['y0']], bottom=[['N'],['R']])) def _(f=input_box(default=(x*y^2)/(x^2+y^4),width=30,label='$f(x)$'), N=slider(5,100,1,10,label='Number of Contours'), R=slider(0.1,1,0.01,1,label='Radius of circular neighborhood'), x0=input_box(0,width=10,label='$x_0$'), y0=input_box(0,width=10,label='$y_0$')): pretty_print(html('Enter $(x_0 ,y_0 )$ above and see what happens as the number of contour levels $\\rightarrow \infty $.')) pretty_print(html('A surface will have a limit in the center of this graph provided there is not a sudden change in color there.')) # Need to make certain the min and max contour lines are not huge due to asymptotes. If so, clip and start contours at some reasonable # values so that there are a nice collection of contours to show around the desired point. surface = contour_plot(f,(x,x0-1,x0+1),(y,y0-1,y0+1),cmap=True,colorbar=True,fill=False,contours=N) surface += parametric_plot([R*cos(u),R*sin(u)],[0,2*pi],color='black') # Nice to use if f=x*y^2/(x^2 + y^4) # var('u') # surface += parametric_plot([u^2,u],[u,-1,1],color='black') limit_point = point((x0,y0),color='red',size=30) # show(limit_point+surface) pretty_print(table([[surface],['hi']])) }}} {{attachment:3D_Limit_Defn_Contours.png}} == Directional Derivatives == This interact displays graphically a tangent line to a function, illustrating a directional derivative (the slope of the tangent line). {{{#!sagecell var('x,y,t,z') f(x,y)=sin(x)*cos(y) pif = float(pi) line_thickness=3 surface_color='blue' plane_color='purple' line_color='red' tangent_color='green' gradient_color='orange' @interact def myfun(location=input_grid(1, 2, default=[0,0], label = "Location (x,y)", width=2), angle=slider(0, 2*pif, label = "Angle"), show_surface=("Show surface", True)): location3d = vector(location[0]+[0]) location = location3d[0:2] direction3d = vector(RDF, [cos(angle), sin(angle), 0]) direction=direction3d[0:2] cos_angle = math.cos(angle) sin_angle = math.sin(angle) df = f.gradient() direction_vector=line3d([location3d, location3d+direction3d], arrow_head=True, rgbcolor=line_color, thickness=line_thickness) curve_point = (location+t*direction).list() curve = parametric_plot(curve_point+[f(*curve_point)], (t,-3,3),color=line_color,thickness=line_thickness) plane = parametric_plot((cos_angle*x+location[0],sin_angle*x+location[1],t), (x, -3,3), (t,-3,3),opacity=0.8, color=plane_color) pt = point3d(location3d.list(),color='green', size=10) tangent_line = parametric_plot((location[0]+t*cos_angle, location[1]+t*sin_angle, f(*location)+t*df(*location)*(direction)), (t, -3,3), thickness=line_thickness, color=tangent_color) picture3d = direction_vector+curve+plane+pt+tangent_line picture2d = contour_plot(f(x,y), (x,-3,3),(y,-3,3), plot_points=100) picture2d += arrow(location.list(), (location+direction).list()) picture2d += point(location.list(),rgbcolor='green',pointsize=40) if show_surface: picture3d += plot3d(f, (x,-3,3),(y,-3,3),opacity=0.7) dff = df(location[0], location[1]) dff3d = vector(RDF,dff.list()+[0]) picture3d += line3d([location3d, location3d+dff3d], arrow_head=True, rgbcolor=gradient_color, thickness=line_thickness) picture2d += arrow(location.list(), (location+dff).list(), rgbcolor=gradient_color, width=line_thickness) show(picture3d,aspect=[1,1,1], axes=True) show(picture2d, aspect_ratio=1) }}} {{attachment:directional derivative.png}} == 3D graph with points and curves == By Robert Marik This sagelet is handy when showing local, constrained and absolute maxima and minima in two variables. Available as a [[http://user.mendelu.cz/marik/sage/3Dgraph_with_points.sws|worksheet]] {{{#!sagecell x,y, t, u, v =var('x y t u v') INI_func='x^2-2*x+y^2-2*y' INI_box='-1,3.2,-1,3.2' INI_points='(1,1,\'green\'),(3/2,3/2),(0,1),(1,0),(0,0,\'black\'),(3,0,\'black\'),(0,3,\'black\')' INI_curves='(t,0,0,3,\'red\'),(0,t,0,3,\'green\'),(t,3-t,0,3)' @interact def _(func=input_box(INI_func,label="f(x,y)=",type=str),\ bounds=input_box(INI_box,label="xmin,xmax,ymin,ymax",type=str),\ st_points=input_box(INI_points,\ label="points <br><small><small>(comma separated pairs, optionally with color)</small></small>", type=str),\ bnd_curves=input_box(INI_curves,label="curves on boundary<br> <small><small><i>(x(t),y(t),tmin,tmax,'opt_color')</i></small></small>", type=str),\ show_planes=("Show zero planes", False), show_axes=("Show axes", True), show_table=("Show table", True)): f=sage_eval('lambda x,y: ' + func) pretty_print(html(r'Function $ f(x,y)=%s$ '%latex(f(x,y)))) xmin,xmax,ymin,ymax=sage_eval('('+bounds+')') A=plot3d(f(x,y),(x,xmin,xmax),(y,ymin,ymax),opacity=0.5) if not(bool(st_points=='')): st_p=sage_eval('('+st_points+',)') html(r'<table border=1>') for current in range(len(st_p)): point_color='red' if bool(len(st_p[current])==3): point_color=st_p[current][2] x0=st_p[current][0] y0=st_p[current][1] z0=f(x0,y0) if show_table: pretty_print(html(r'<tr><td>$\quad f(%s,%s)\quad $</td><td>$\quad %s$</td>\ </tr>'%(latex(x0),latex(y0),z0.n()))) A=A+point3d((x0,y0,z0),size=9,rgbcolor=point_color) html(r'</table>') if not(bool(bnd_curves=='')): bnd_cc=sage_eval('('+bnd_curves+',)',locals={'t':t}) for current in range(len(bnd_cc)): bnd_c=bnd_cc[current]+('black',) A=A+parametric_plot3d((bnd_c[0],bnd_c[1],f(bnd_c[0],bnd_c[1])),\ (t,bnd_c[2],bnd_c[3]),thickness=3,rgbcolor=bnd_c[4]) if show_planes: A=A+plot3d(0,(x,xmin,xmax),(y,ymin,ymax),opacity=0.3,rgbcolor='gray') zmax=A.bounding_box()[1][2] zmin=A.bounding_box()[0][2] A=A+parametric_plot3d((u,0,v),(u,xmin,xmax),(v,zmin,zmax),opacity=0.3,rgbcolor='gray') A=A+parametric_plot3d((0,u,v),(u,ymin,ymax),(v,zmin,zmax),opacity=0.3,rgbcolor='gray') if show_axes: zmax=A.bounding_box()[1][2] zmin=A.bounding_box()[0][2] A=A+line3d([(xmin,0,0), (xmax,0,0)], arrow_head=True,rgbcolor='black') A=A+line3d([(0,ymin,0), (0,ymax,0)], arrow_head=True,rgbcolor='black') A=A+line3d([(0,0,zmin), (0,0,zmax)], arrow_head=True,rgbcolor='black') show(A) }}} {{attachment:3Dgraph_with_points.png}} == Approximating function in two variables by differential == by Robert Marik {{{#!sagecell x,y=var('x y') html('<h2>Explaining approximation of a function in two \ variables by differential</h2>') pretty_print(html('Points x0 and y0 are values where the exact value of the function \ is known. Deltax and Deltay are displacements of the new point. Exact value \ and approximation by differential at shifted point are compared.')) @interact def _(func=input_box('sqrt(x^3+y^3)',label="f(x,y)=",type=str), x0=1, y0=2, \ deltax=slider(-1,1,0.01,0.2),\ deltay=slider(-1,1,0.01,-0.4), xmin=0, xmax=2, ymin=0, ymax=3): f=sage_eval('lambda x,y: ' + func) derx(x,y)=diff(f(x,y),x) dery(x,y)=diff(f(x,y),y) tangent(x,y)=f(x0,y0)+derx(x0,y0)*(x-x0)+dery(x0,y0)*(y-y0) A=plot3d(f(x,y),(x,xmin,xmax),(y,ymin,ymax),opacity=0.5) B=plot3d(tangent(x,y),(x,xmin,xmax),(y,ymin,ymax),color='red',opacity=0.5) C=point3d((x0,y0,f(x0,y0)),rgbcolor='blue',size=9) CC=point3d((x0+deltax,y0+deltay,f(x0+deltax,y0+deltay)),rgbcolor='blue',size=9) D=point3d((x0+deltax,y0+deltay,tangent(x0+deltax,y0+deltay)),rgbcolor='red',size=9) exact_value_ori=f(x0,y0).n(digits=10) exact_value=f(x0+deltax,y0+deltay) approx_value=tangent(x0+deltax,y0+deltay).n(digits=10) abs_error=(abs(exact_value-approx_value)) pretty_print(html(r'Function $ f(x,y)=%s \approx %s $ '%(latex(f(x,y)),latex(tangent(x,y))))) pretty_print(html(r' $f %s = %s$'%(latex((x0,y0)),latex(exact_value_ori)))) pretty_print(html(r'Shifted point $%s$'%latex(((x0+deltax),(y0+deltay))))) pretty_print(html(r'Value of the function in shifted point is $%s$'%f(x0+deltax,y0+deltay))) pretty_print(html(r'Value on the tangent plane in shifted point is $%s$'%latex(approx_value))) pretty_print(html(r'Error is $%s$'%latex(abs_error))) show(A+B+C+CC+D) }}} {{attachment:3D_differential.png}} == Taylor approximations in two variables == by John Palmieri This displays the nth order Taylor approximation, for n from 1 to 10, of the function sin(x^2^ + y^2^) cos(y) exp(-(x^2^+y^2^)/2). {{{#!sagecell var('x y') var('xx yy') G = sin(xx^2 + yy^2) * cos(yy) * exp(-0.5*(xx^2+yy^2)) def F(x,y): return G.subs(xx=x).subs(yy=y) plotF = plot3d(F, (0.4, 2), (0.4, 2), adaptive=True, color='blue') @interact def _(x0=(0.5,1.5), y0=(0.5, 1.5), order=[1..10]): F0 = float(G.subs(xx=x0).subs(yy=y0)) P = (x0, y0, F0) dot = point3d(P, size=15, color='red') plot = dot + plotF approx = F0 for n in range(1, order+1): for i in range(n+1): if i == 0: deriv = G.diff(yy, n) elif i == n: deriv = G.diff(xx, n) else: deriv = G.diff(xx, i).diff(yy, n-i) deriv = float(deriv.subs(xx=x0).subs(yy=y0)) coeff = binomial(n, i)/factorial(n) approx += coeff * deriv * (x-x0)^i * (y-y0)^(n-i) plot += plot3d(approx, (x, 0.4, 1.6), (y, 0.4, 1.6), color='red', opacity=0.7) pretty_print(html('$F(x,y) = e^{-(x^2+y^2)/2} \\cos(y) \\sin(x^2+y^2)$')) show(plot) }}} {{attachment:taylor-3d.png}} == Volumes over non-rectangular domains == by John Travis https://cloud.sagemath.com/projects/19575ea0-317e-402b-be57-368d04c113db/files/pub/2801-2901/2829.sagews {{{#!sagecell ## Graphing surfaces over non-rectangular domains ## John Travis ## Spring 2011 ## ## ## An updated version of this worksheet may be available at http://sagenb.mc.edu ## ## Interact allows the user to input up to two inequality constraints on the ## domain when dealing with functional surfaces ## ## User inputs: ## f = "top" surface with z = f(x,y) ## g = "bottom" surface with z = g(x,y) ## condition1 = a single boundary constraint. It should not include && or | to join two conditions. ## condition2 = another boundary constraint. If there is only one constraint, just enter something true ## or even just an x (or y) in the entry blank. ## ## var('x,y') # f is the top surface # g is the bottom surface global f,g # condition1 and condition2 are the inequality constraints. It would be nice # to have any number of conditions connected by $$ or | global condition1,condition2 @interact def _(f=input_box(default=(1/3)*x^2 + (1/4)*y^2 + 5,label='$f(x)=$'), g=input_box(default=-1*x+0*y,label='$g(x)=$'), condition1=input_box(default= x^2+y^2<8,label='$Constraint_1=$'), condition2=input_box(default=y<sin(3*x),label='$Constraint_2=$'), show_3d=('Stereographic',false), show_vol=('Shade volume',true), dospin = ('Spin?',true), clr = color_selector('#faff00', label='Volume Color', widget='colorpicker', hide_box=True), xx = range_slider(-5, 5, 1, default=(-3,3), label='X Range'), yy = range_slider(-5, 5, 1, default=(-3,3), label='Y Range'), auto_update=false): # This is the top function actually graphed by using NaN outside domain def F(x,y): if condition1(x=x,y=y): if condition2(x=x,y=y): return f(x=x,y=y) else: return -NaN else: return -NaN # This is the bottom function actually graphed by using NaN outside domain def G(x,y): if condition1(x=x,y=y): if condition2(x=x,y=y): return g(x=x,y=y) else: return -NaN else: return -NaN P = Graphics() # The graph of the top and bottom surfaces P_list = [] P_list.append(plot3d(F,(x,xx[0],xx[1]),(y,yy[0],yy[1]),color='blue',opacity=0.9)) P_list.append(plot3d(G,(x,xx[0],xx[1]),(y,yy[0],yy[1]),color='gray',opacity=0.9)) # Interpolate "layers" between the top and bottom if desired if show_vol: ratios = range(10) def H(x,y,r): return (1-r)*F(x=x,y=y)+r*G(x=x,y=y) P_list.extend([ plot3d(lambda x,y: H(x,y,ratios[1]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[2]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[3]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[4]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[5]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[6]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[7]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[8]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr), plot3d(lambda x,y: H(x,y,ratios[9]/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2,color=clr) ]) # P = plot3d(lambda x,y: H(x,y,ratio/10),(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.1) # Now, accumulate all of the graphs into one grouped graph. P = sum(P_list[i] for i in range(len(P_list))) if show_3d: show(P,frame=true,axes=false,xmin=xx[0],xmax=xx[1],ymin=yy[0],ymax=yy[1],stereo='redcyan',figsize=(6,9),viewer='jmol',spin=dospin) else: show(P,frame=true,axes=false,xmin=xx[0],xmax=xx[1],ymin=yy[0],ymax=yy[1],figsize=(6,9),viewer='jmol',spin=dospin) }}} {{attachment:3D_Irregular_Volume.png}} == Lateral Surface Area (FIXME in Jupyter) == by John Travis http://sagenb.mc.edu/home/pub/89/ {{{#!sagecell ## Display and compute the area of the lateral surface between two surfaces ## corresponding to the (scalar) line integral ## John Travis ## Spring 2011 ## var('x,y,t,s') @interact(layout=dict(top=[['f','u'],['g','v']], left=[['a'],['b'],['in_3d'],['smoother']], bottom=[['xx','yy']])) def _(f=input_box(default=6-4*x^2-y^2*2/5,label='Top = $f(x,y) = $',width=30), g=input_box(default=-2+sin(x)+sin(y),label='Bottom = $g(x,y) = $',width=30), u=input_box(default=cos(t),label=' $ x = u(t) = $',width=20), v=input_box(default=2*sin(t),label=' $ y = v(t) = $',width=20), a=input_box(default=0,label='$a = $',width=10), b=input_box(default=3*pi/2,label='$b = $',width=10), xx = range_slider(-5, 5, 1, default=(-1,1), label='x view'), yy = range_slider(-5, 5, 1, default=(-2,2), label='y view'), in_3d = checkbox(default=true,label='3D'), smoother=checkbox(default=false), auto_update=true): ds = sqrt(derivative(u,t)^2+derivative(v,t)^2) # Set up the integrand to compute the line integral, making all attempts # to simplify the result so that it looks as nice as possible. A = (f(x=u,y=v)-g(x=u,y=v))*ds.simplify_trig().simplify() # It is not expected that Sage can actually perform the line integral calculation. # So, the result displayed may not be a numerical value as expected. # Creating a good but harder example that "works" is desirable. # If you want Sage to try, uncomment the lines below. # line_integral = integrate(A,t,a,b) # html(r'<align=center size=+1>Lateral Surface Area = $ %s $ </font>'%latex(line_integral)) line_integral_approx = numerical_integral(A,a,b)[0] pretty_print(html(r'<font align=center size=+1>Lateral Surface $ \approx $ %s</font>'%str(line_integral_approx))) # Plot the top function z = f(x,y) that is being integrated. G = plot3d(f,(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2) G += plot3d(g,(x,xx[0],xx[1]),(y,yy[0],yy[1]),opacity=0.2) # Add space curves on the surfaces "above" the domain curve (u(t),v(t)) G += parametric_plot3d([u,v,g(x=u,y=v)],(t,a,b),thickness=2,color='red') G += parametric_plot3d([u,v,f(x=u,y=v)],(t,a,b),thickness=2,color='red') k=0 if smoother: delw = 0.025 lat_thick = 3 else: delw = 0.10 lat_thick = 10 for w in (a,a+delw,..,b): G += parametric_plot3d([u(t=w),v(t=w),s*f(x=u(t=w),y=v(t=w))+(1-s)*g(x=u(t=w),y=v(t=w))],(s,0,1),thickness=lat_thick,color='yellow',opacity=0.9) if in_3d: show(G,stereo='redcyan',spin=true) else: show(G,perspective_depth=true,spin=true) }}} {{attachment:Lateral_Surface.png}} == Parametric surface example (FIXME in Jupyter) == by Marshall Hampton {{{#!sagecell var('u,v') npi = RDF(pi) @interact def viewer(mesh = checkbox(default = False, label = 'Show u,v meshlines'), uc = slider(-2,2,1/10,0, label = '<span style="color:red">Constant u value</span>'), vc = slider(-2,2,1/10,0, label = '<span style="color:green">Constant v value</span>'), functions = input_box([u,v^2,u^2+v])): f1(u,v) = functions[0] f2(u,v) = functions[1] f3(u,v) = functions[2] surface_plot = parametric_plot3d([f1,f2,f3], (u,-2,2), (v,-2,2), mesh = mesh, opacity = .8) constant_u = line3d([[f1(uc,q), f2(uc,q), f3(uc,q)] for q in srange(-2,2,.01)], rgbcolor = (1,0,0), thickness = 3) constant_v = line3d([[f1(q,vc), f2(q,vc), f3(q,vc)] for q in srange(-2,2,.01)], rgbcolor = (0,1,0), thickness = 3) show(surface_plot + constant_u + constant_v, frame = False) }}} {{attachment:parametric_surface.png}} == Line Integrals in 3D Vector Field == by John Travis https://cloud.sagemath.com/projects/19575ea0-317e-402b-be57-368d04c113db/files/pub/2801-2901/2827-$%20%5Cint_%7BC%7D%20%5Cleft%20%5Clangle%20M,N,P%20%5Cright%20%5Crangle%20dr%20$%20=%20$%20%25s%20$.sagews {{{#!sagecell ## This worksheet interactively computes and displays the line integral of a 3D vector field ## over a given smooth curve C ## ## John Travis ## Mississippi College ## 06/16/11 ## ## An updated version of this worksheet may be available at http://sagenb.mc.edu ## var('x,y,z,t,s') @interact def _(M=input_box(default=x*y*z,label="$M(x,y,z)$"), N=input_box(default=-y*z,label="$N(x,y,z)$"), P=input_box(default=z*y,label="$P(x,y,z)$"), u=input_box(default=cos(t),label="$x=u(t)$"), v=input_box(default=2*sin(t),label="$y=v(t)$"), w=input_box(default=t*(t-2*pi)/pi,label="$z=w(t)$"), tt = range_slider(-2*pi, 2*pi, pi/6, default=(0,2*pi), label='t Range'), xx = range_slider(-5, 5, 1, default=(-1,1), label='x Range'), yy = range_slider(-5, 5, 1, default=(-2,2), label='y Range'), zz = range_slider(-5, 5, 1, default=(-3,1), label='z Range'), in_3d=checkbox(true)): # setup the parts and then compute the line integral dr = [derivative(u(t),t),derivative(v(t),t),derivative(w(t),t)] A = (M(x=u(t),y=v(t),z=w(t))*dr[0] +N(x=u(t),y=v(t),z=w(t))*dr[1] +P(x=u(t),y=v(t),z=w(t))*dr[2]) global line_integral line_integral = integral(A(t=t),t,tt[0],tt[1]) pretty_print(html(r'<h2 align=center>$ \int_{C} \left \langle M,N,P \right \rangle dr $ = $ %s $ </h2>'%latex(line_integral))) G = plot_vector_field3d((M,N,P),(x,xx[0],xx[1]),(y,yy[0],yy[1]),(z,zz[0],zz[1]),plot_points=6) G += parametric_plot3d([u,v,w],(t,tt[0],tt[1]),thickness='5',color='yellow') if in_3d: show(G,stereo='redcyan',spin=true) else: show(G,perspective_depth=true) }}} {{attachment:3D_Line_Integral.png}} |
Sage Interactions - Calculus
goto interact main page
Contents
-
Sage Interactions - Calculus
- Root Finding Using Bisection
- Newton's Method
- A contour map and 3d plot of two inverse distance functions
- A simple tangent line grapher
- Numerical integrals with the midpoint rule
- Numerical integrals with various rules
- Some polar parametric curves
- Function tool
- Newton-Raphson Root Finding
- Coordinate Transformations (FIXME in Jupyter)
- Taylor Series
- Illustration of the precise definition of a limit
- A graphical illustration of sin(x)/x -> 1 as x-> 0
- Quadric Surface Plotter
- The midpoint rule for numerically integrating a function of two variables
- Gaussian (Legendre) quadrature
- Vector Calculus, 2-D Motion
- Vector Calculus, 3-D Motion
- Multivariate Limits by Definition
- Directional Derivatives
- 3D graph with points and curves
- Approximating function in two variables by differential
- Taylor approximations in two variables
- Volumes over non-rectangular domains
- Lateral Surface Area (FIXME in Jupyter)
- Parametric surface example (FIXME in Jupyter)
- Line Integrals in 3D Vector Field
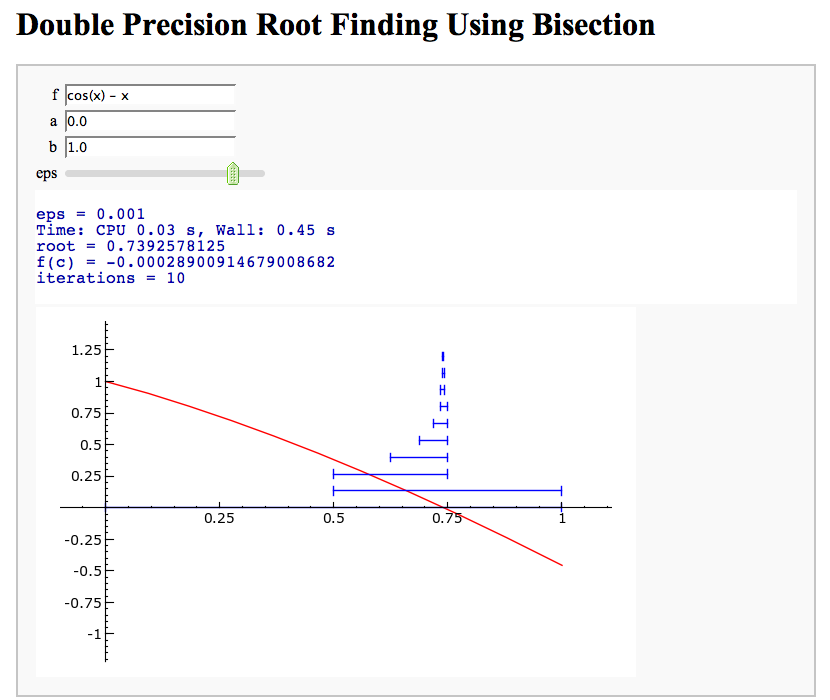
Root Finding Using Bisection
by William Stein
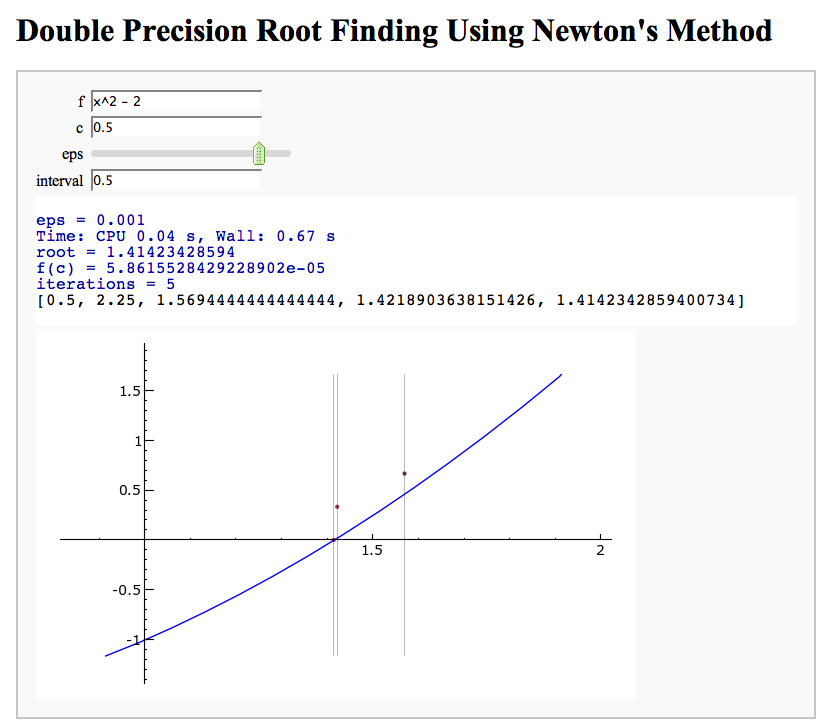
Newton's Method
Note that there is a more complicated Newton's method below.
by William Stein
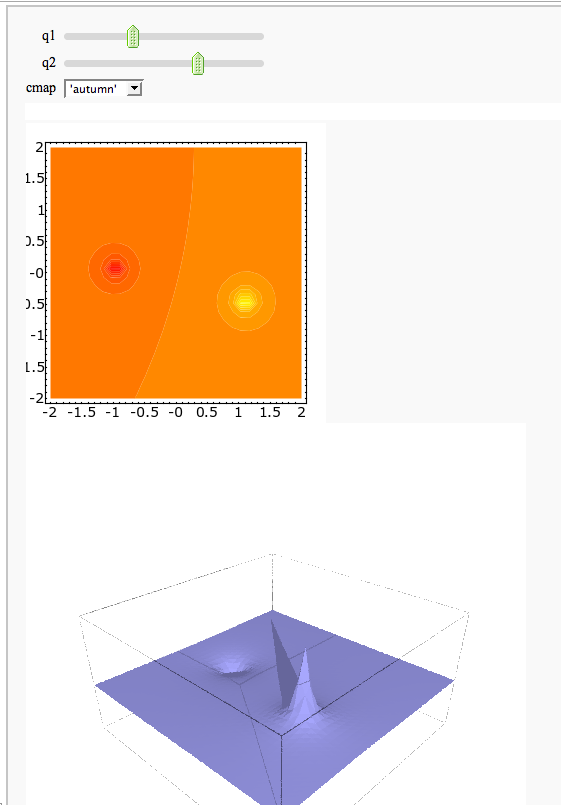
A contour map and 3d plot of two inverse distance functions
by William Stein
A simple tangent line grapher
by Marshall Hampton
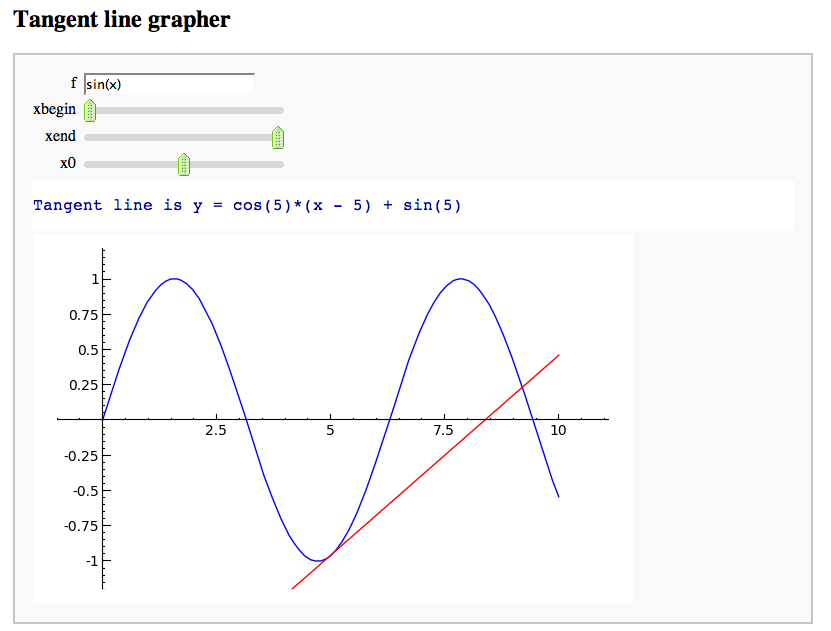
Numerical integrals with the midpoint rule
by Marshall Hampton
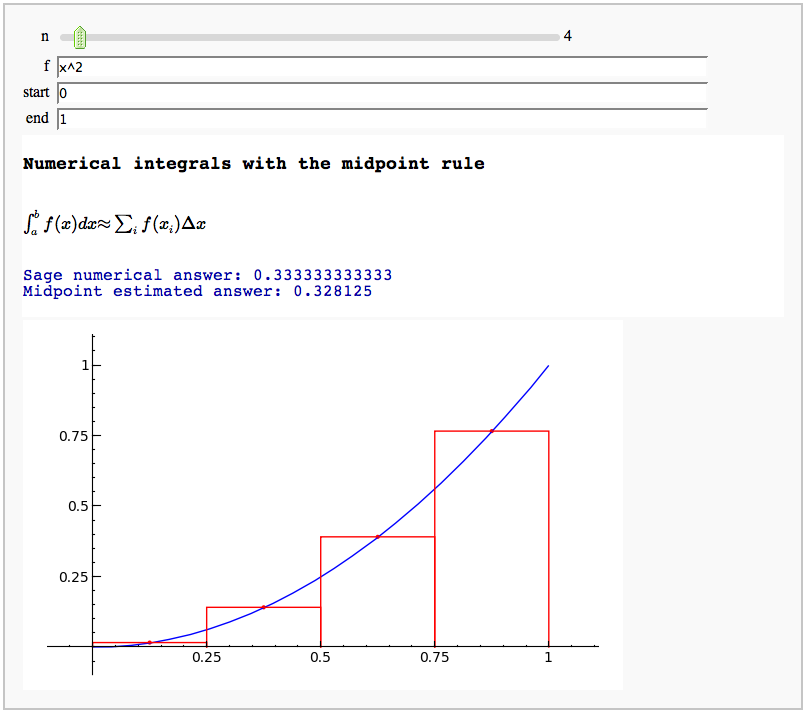
Numerical integrals with various rules
by Nick Alexander (based on the work of Marshall Hampton)
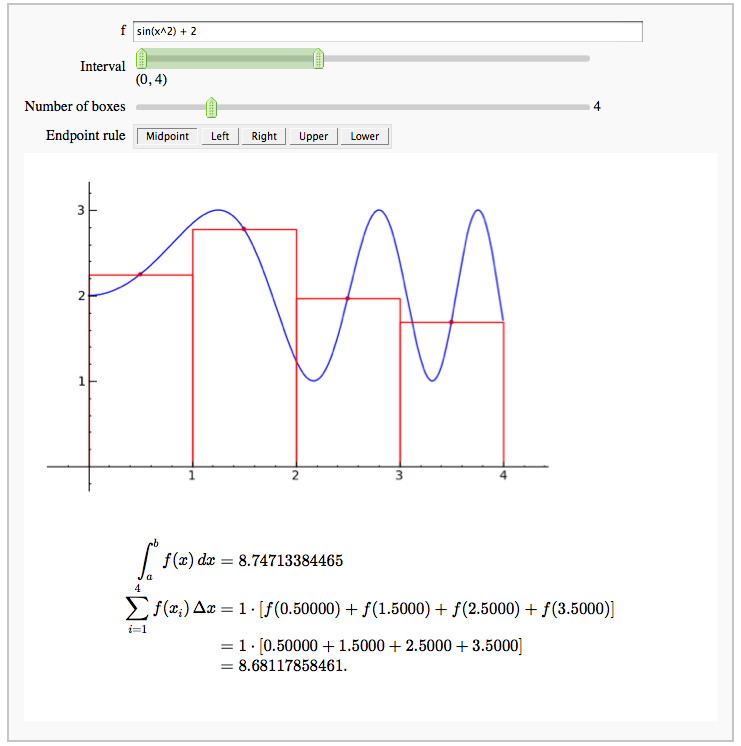
Some polar parametric curves
by Marshall Hampton. This is not very general, but could be modified to show other families of polar curves.
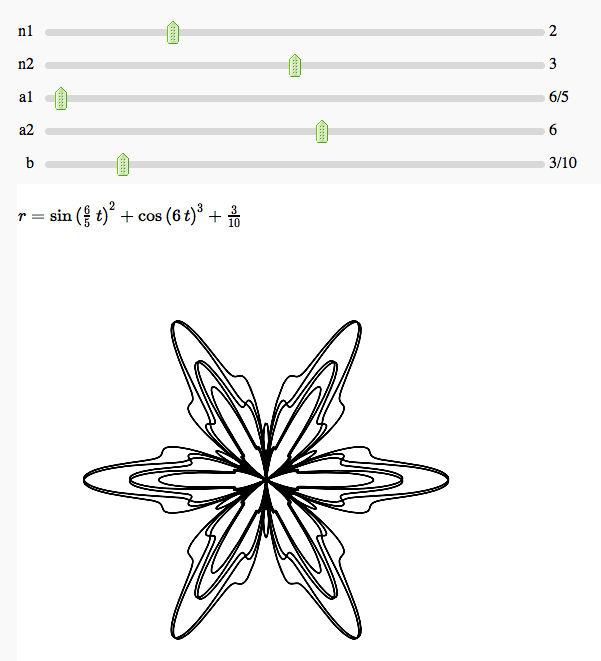
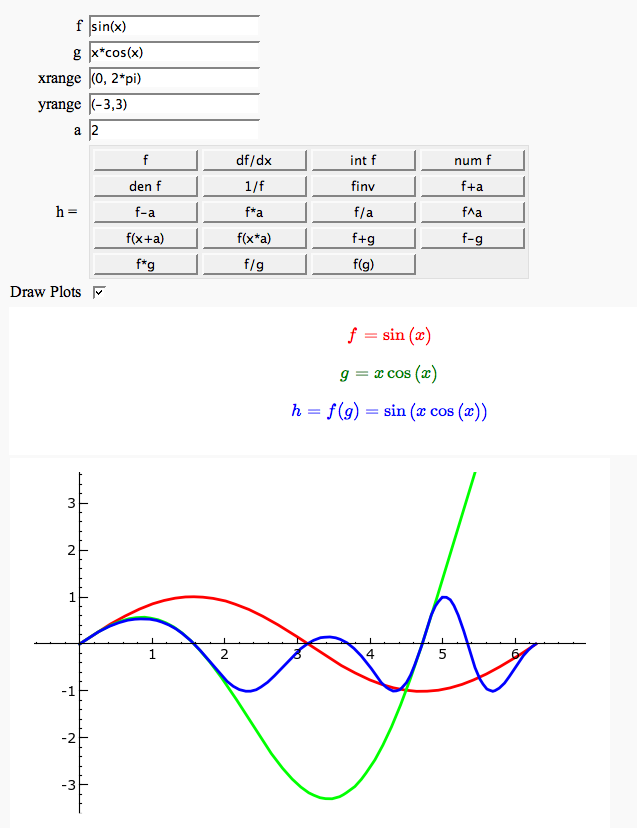
Function tool
Enter symbolic functions f, g, and a, a range, then click the appropriate button to compute and plot some combination of f, g, and a along with f and g. This is inspired by the Matlab funtool GUI.
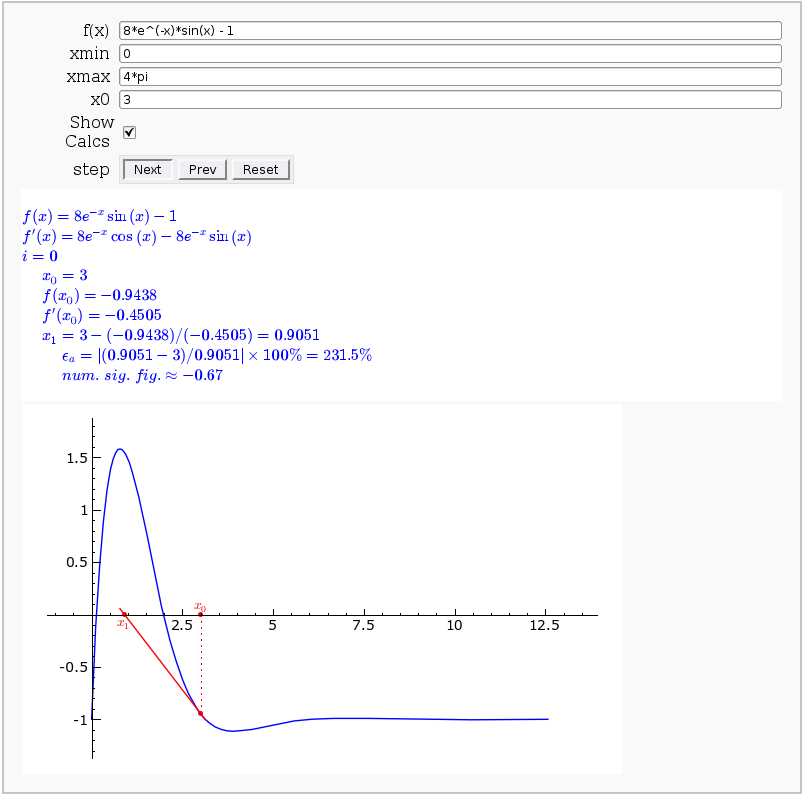
Newton-Raphson Root Finding
by Neal Holtz
This allows user to display the Newton-Raphson procedure one step at a time. It uses the heuristic that, if any of the values of the controls change, then the procedure should be re-started, else it should be continued.
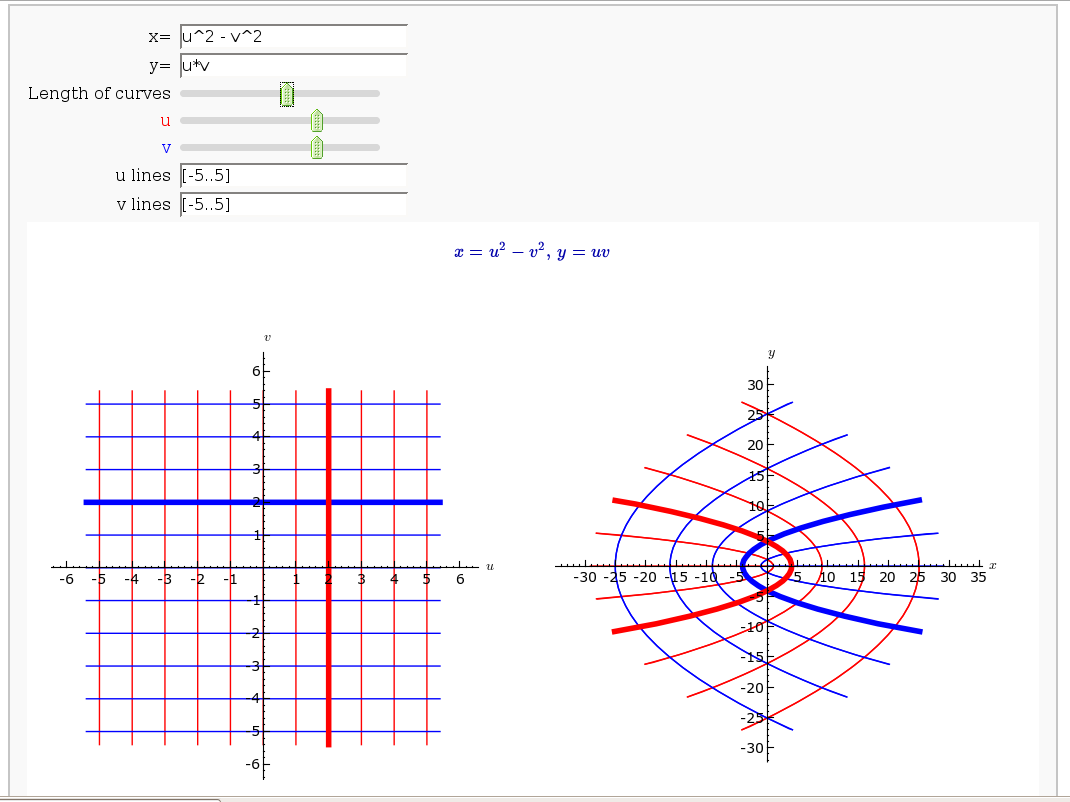
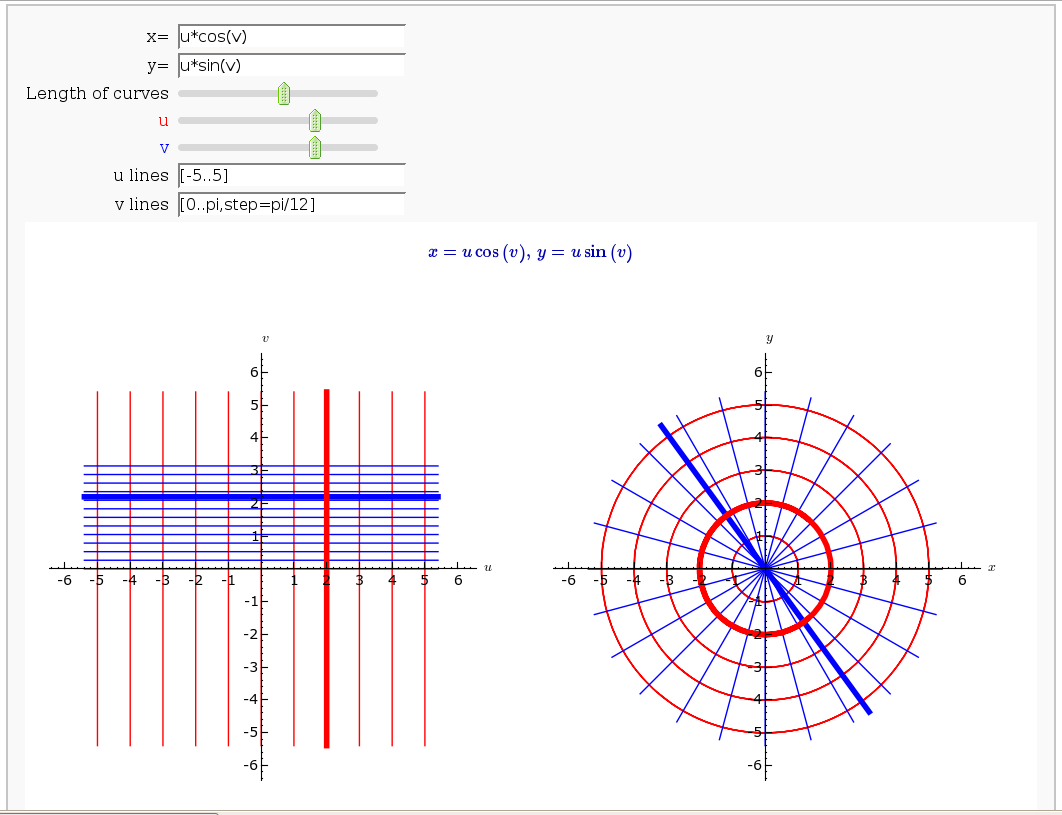
Coordinate Transformations (FIXME in Jupyter)
by Jason Grout
Taylor Series
by Harald Schilly
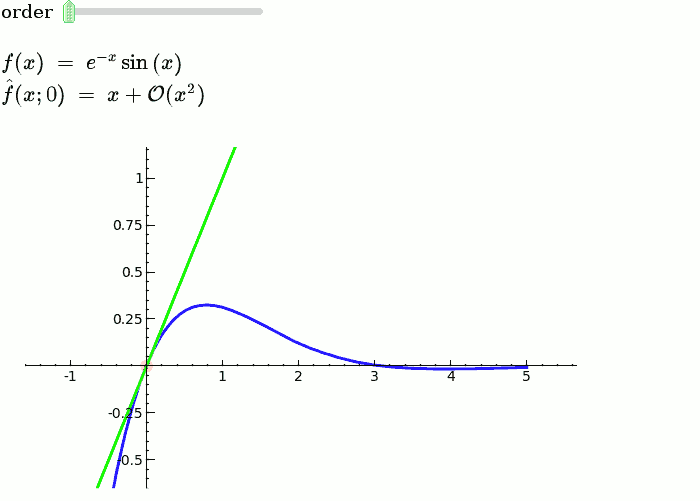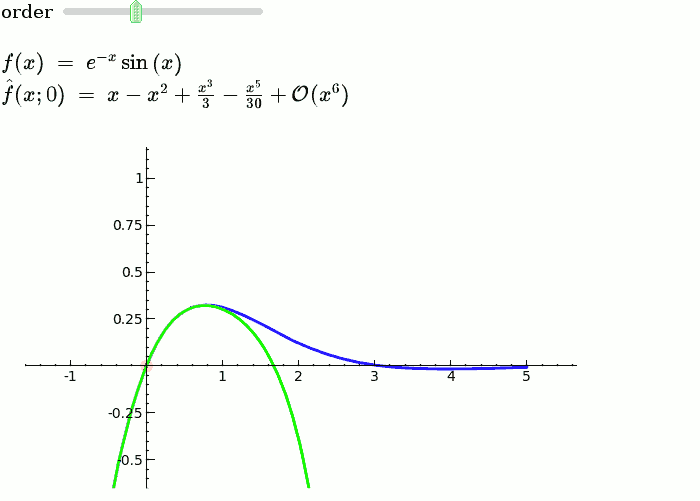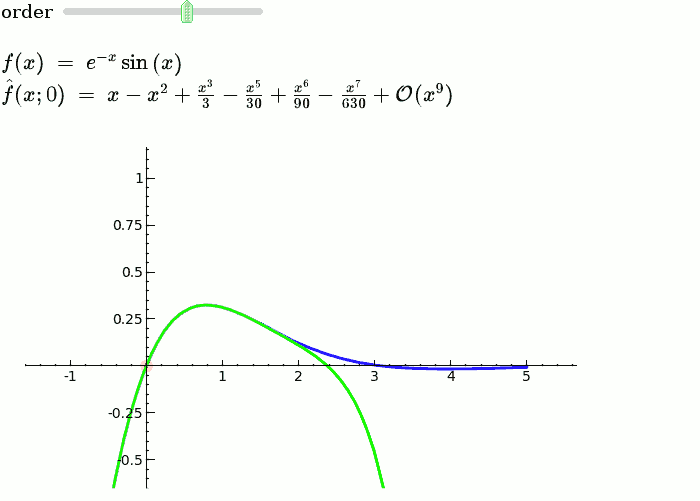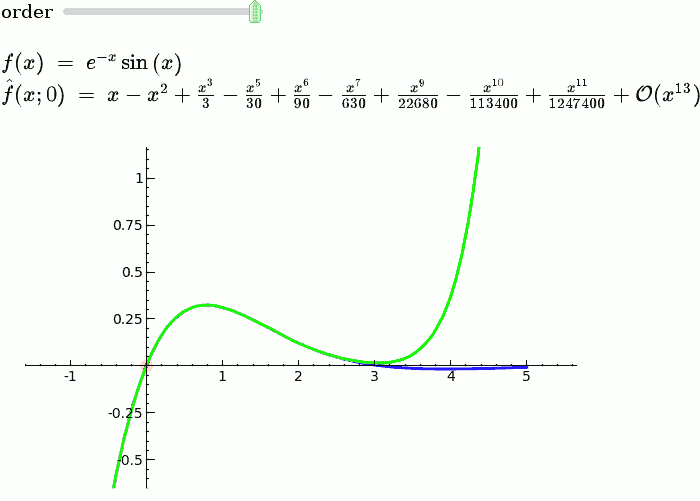
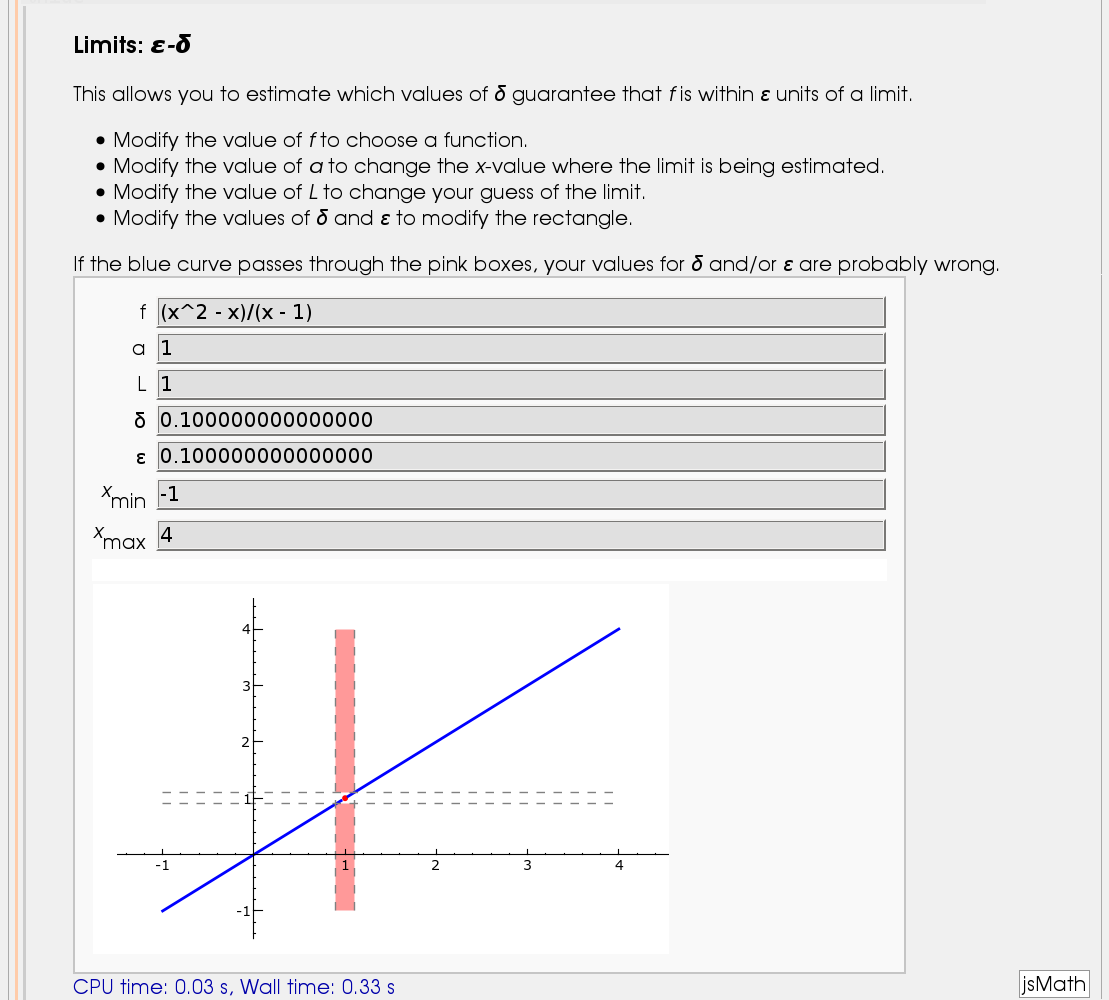
Illustration of the precise definition of a limit
by John Perry
I'll break tradition and put the image first. Apologies if this is Not A Good Thing.
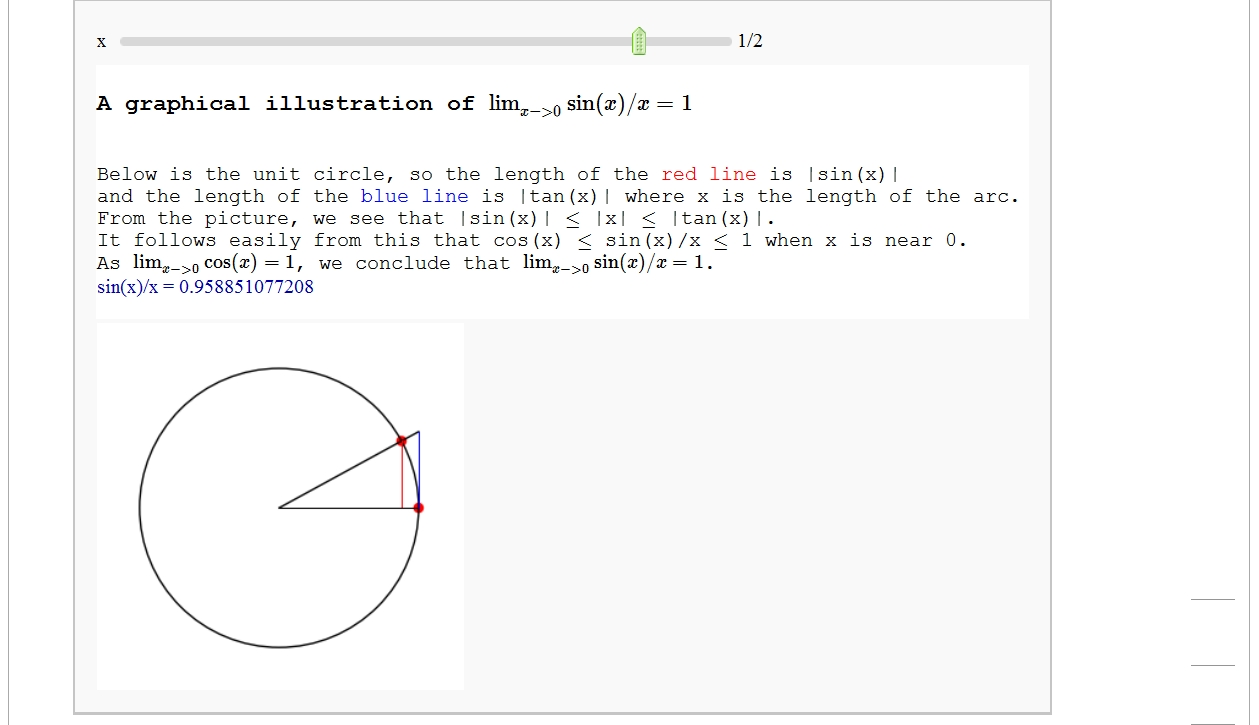
A graphical illustration of sin(x)/x -> 1 as x-> 0
by Wai Yan Pong
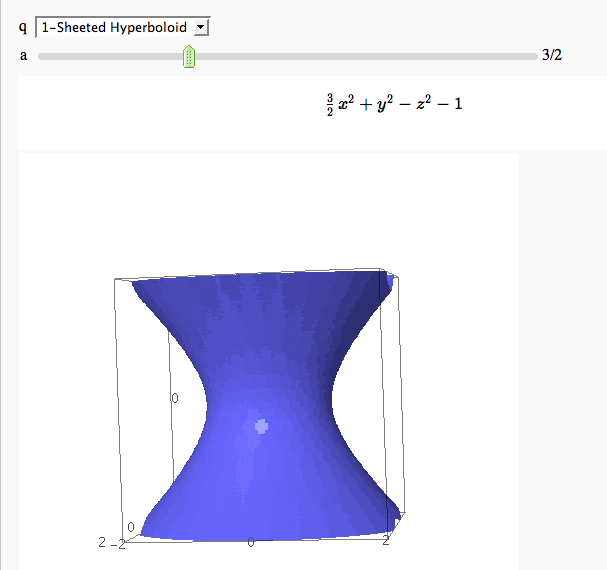
Quadric Surface Plotter
by Marshall Hampton. This is pretty simple, so I encourage people to spruce it up. In particular, it isn't set up to show all possible types of quadrics.
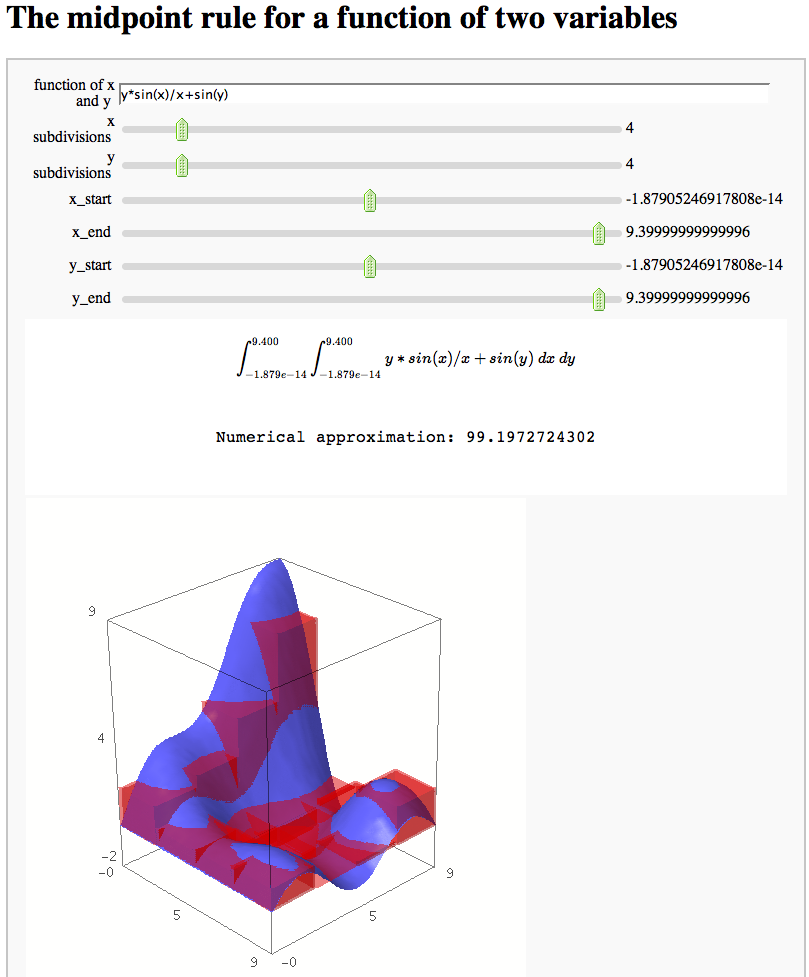
The midpoint rule for numerically integrating a function of two variables
by Marshall Hampton
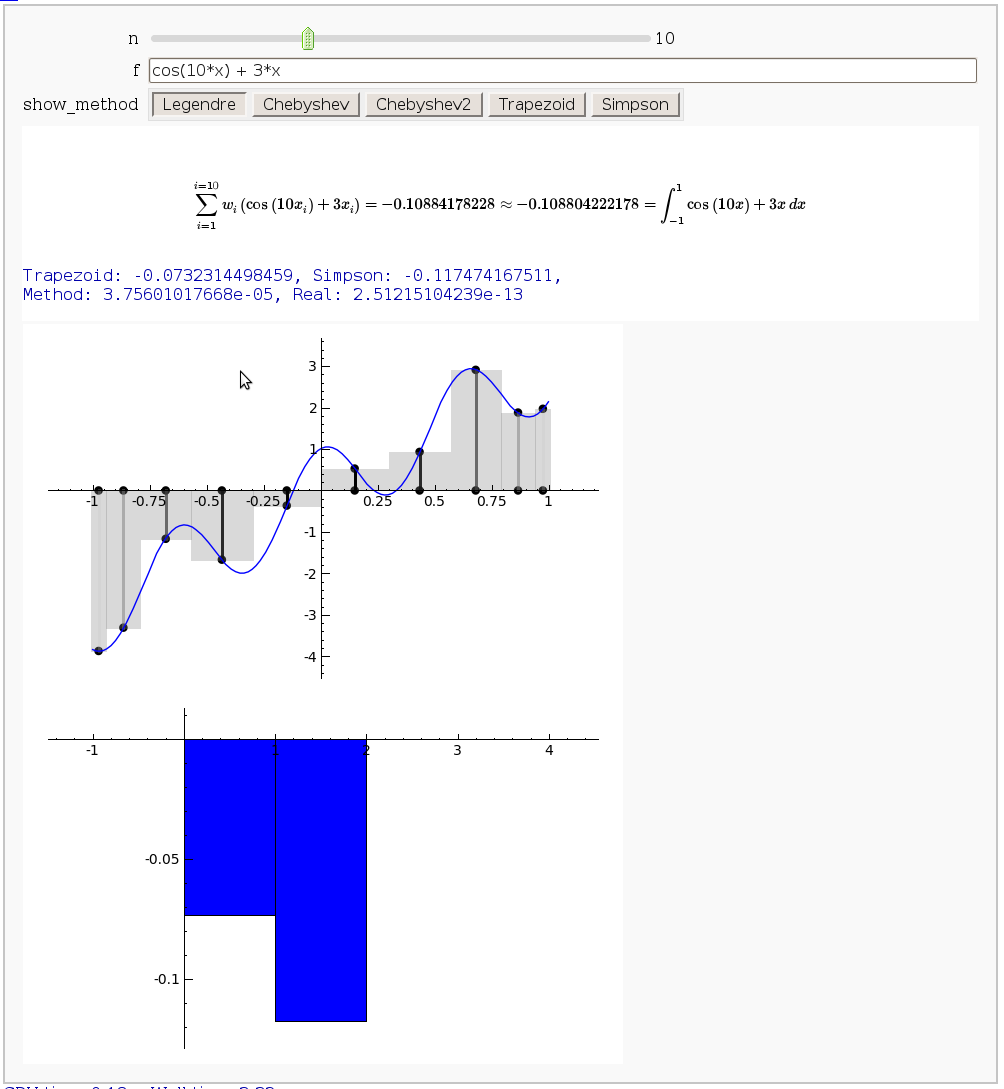
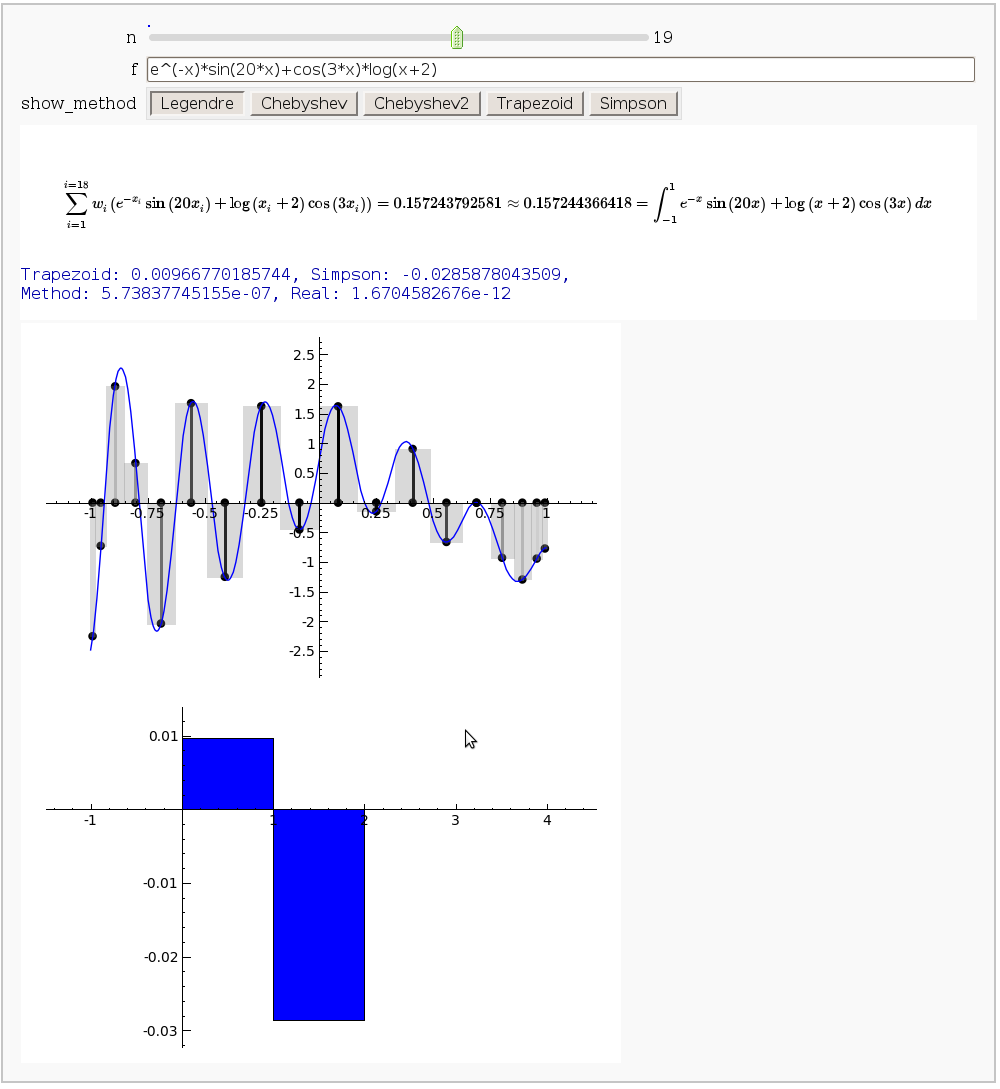
Gaussian (Legendre) quadrature
by Jason Grout
The output shows the points evaluated using Gaussian quadrature (using a weight of 1, so using Legendre polynomials). The vertical bars are shaded to represent the relative weights of the points (darker = more weight). The error in the trapezoid, Simpson, and quadrature methods is both printed out and compared through a bar graph. The "Real" error is the error returned from scipy on the definite integral.
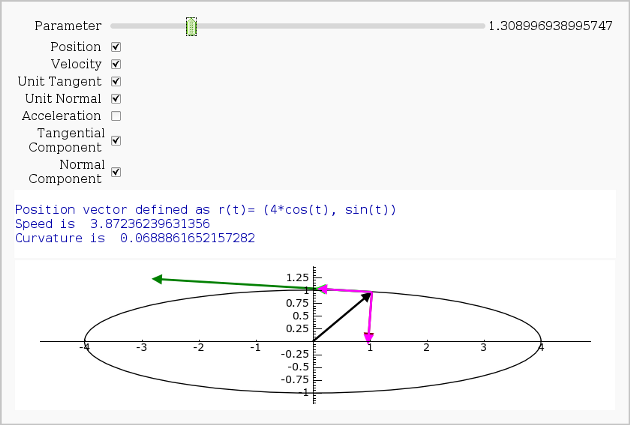
Vector Calculus, 2-D Motion
By Rob Beezer
A fast_float() version is available in a worksheet
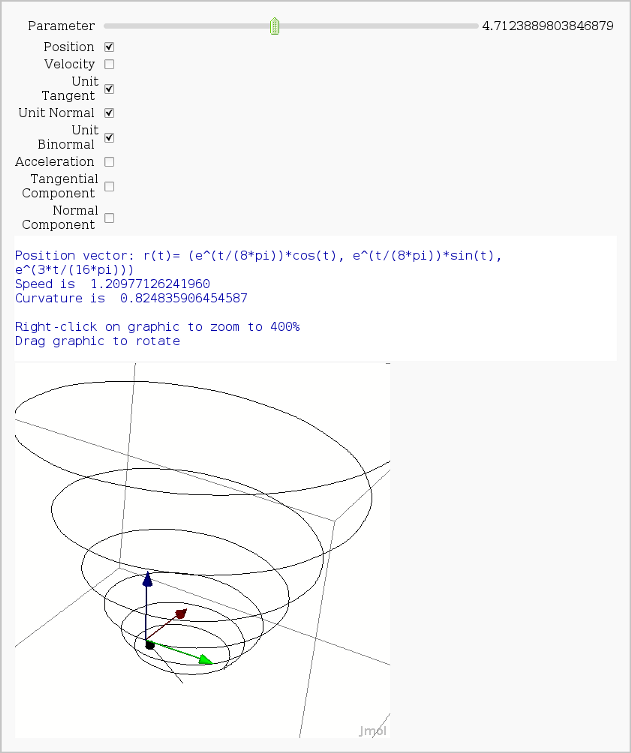
Vector Calculus, 3-D Motion
by Rob Beezer
Available as a worksheet
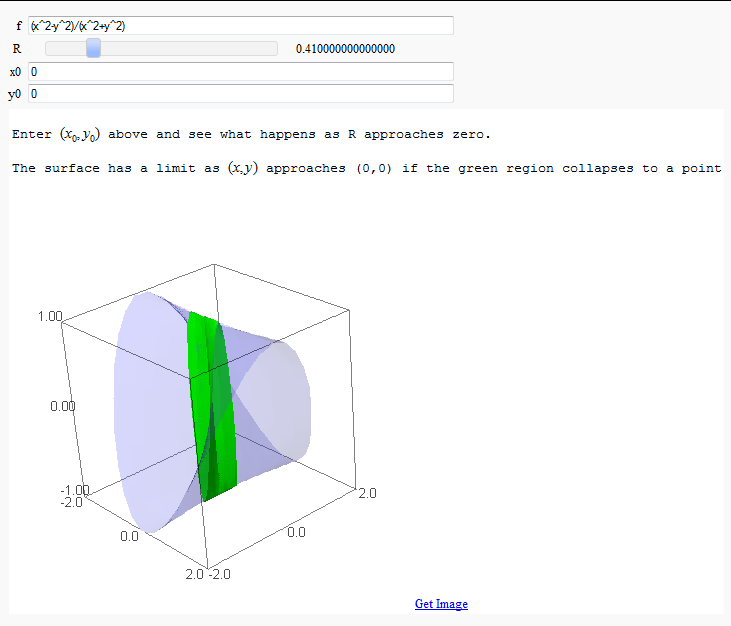
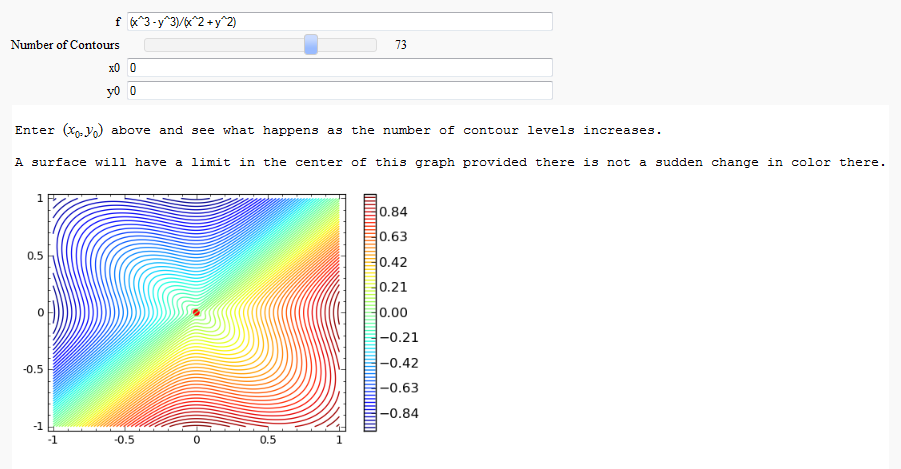
Multivariate Limits by Definition
by John Travis
http://sagenb.mc.edu/home/pub/97/
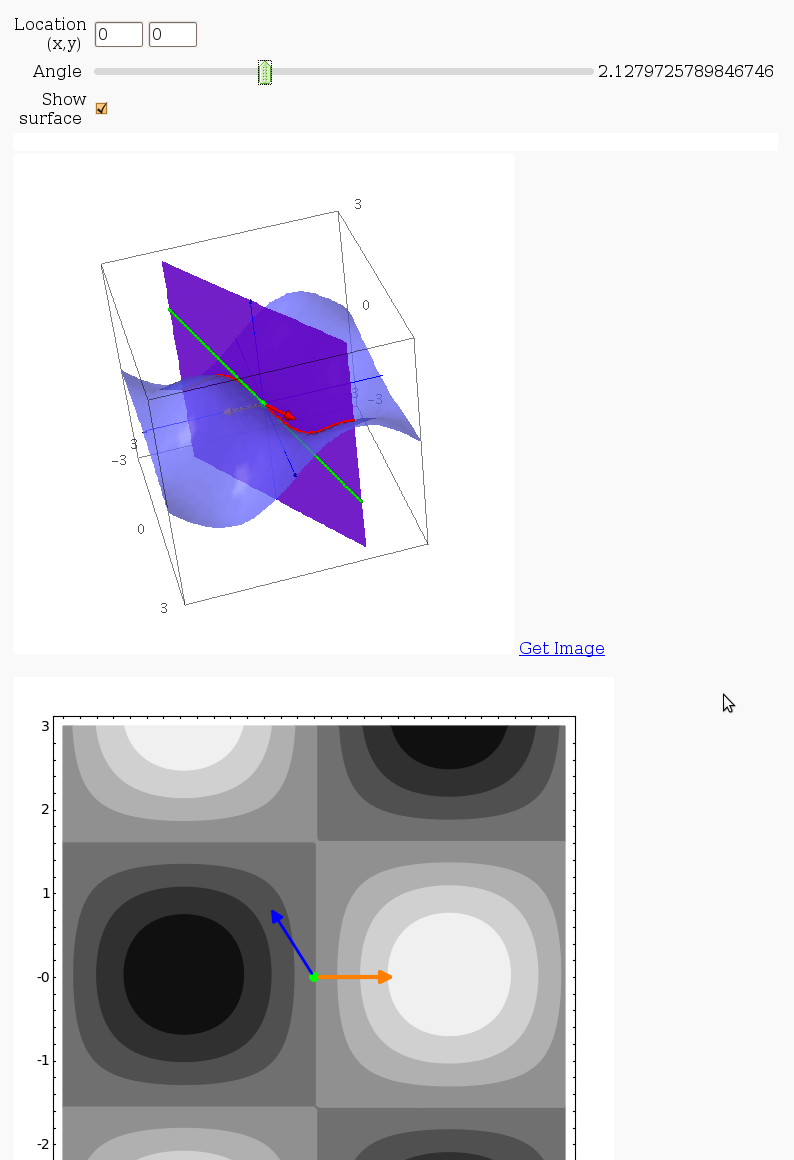
Directional Derivatives
This interact displays graphically a tangent line to a function, illustrating a directional derivative (the slope of the tangent line).
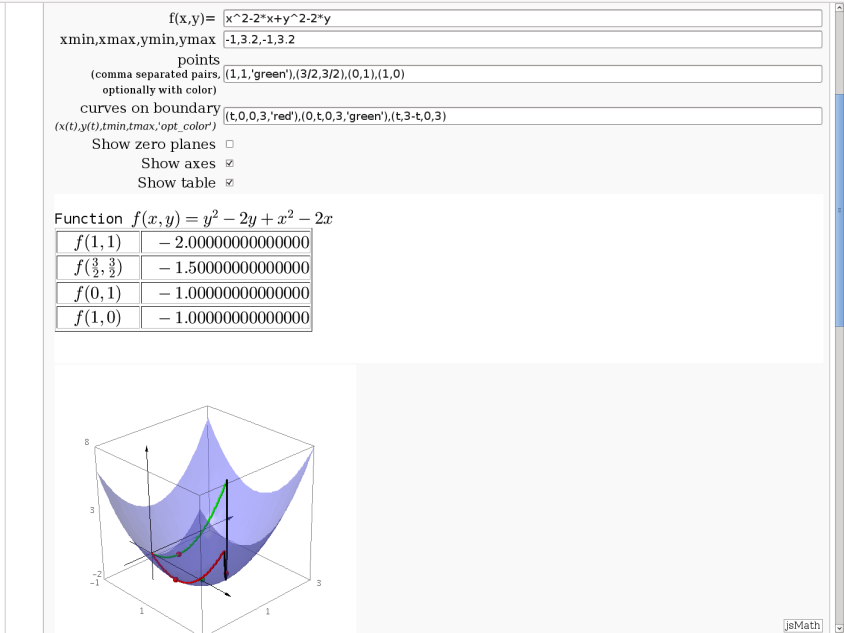
3D graph with points and curves
By Robert Marik
This sagelet is handy when showing local, constrained and absolute maxima and minima in two variables. Available as a worksheet
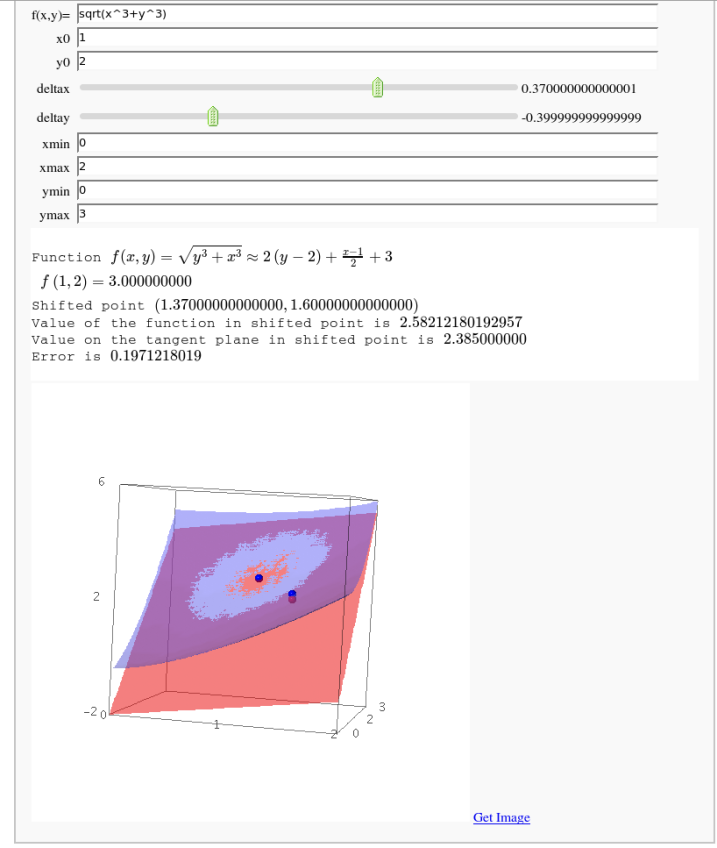
Approximating function in two variables by differential
by Robert Marik
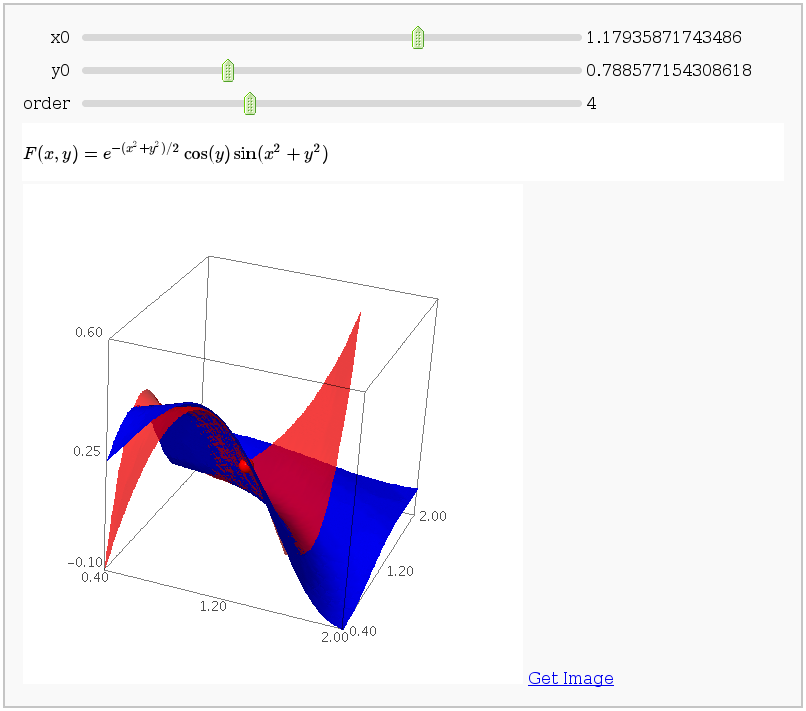
Taylor approximations in two variables
by John Palmieri
This displays the nth order Taylor approximation, for n from 1 to 10, of the function sin(x2 + y2) cos(y) exp(-(x2+y2)/2).
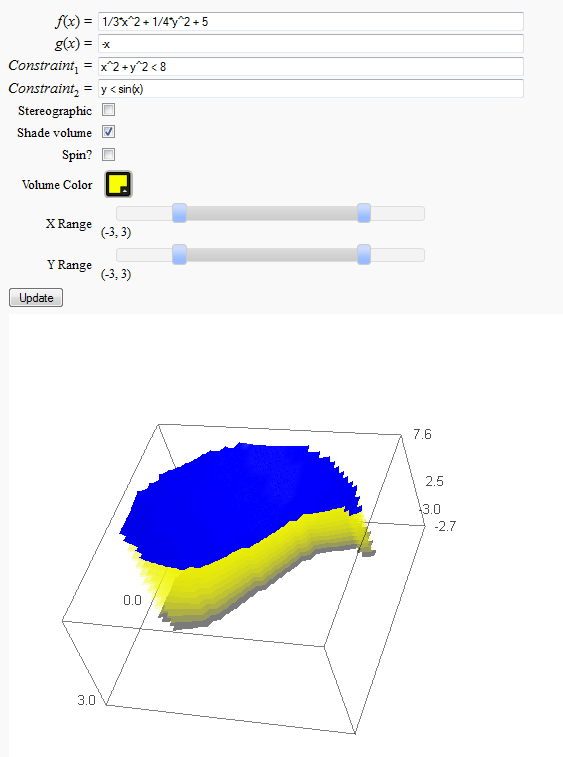
Volumes over non-rectangular domains
by John Travis
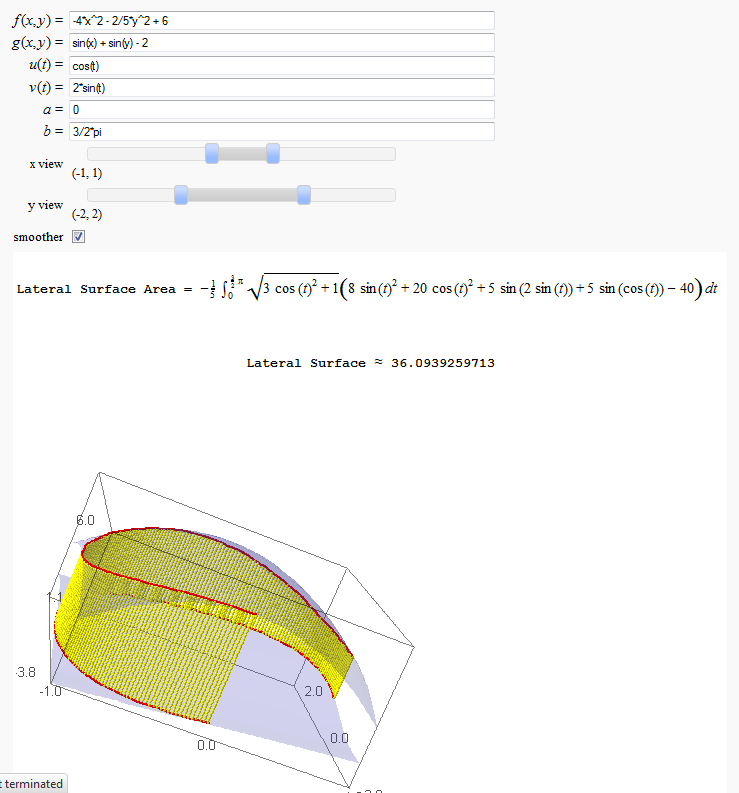
Lateral Surface Area (FIXME in Jupyter)
by John Travis
http://sagenb.mc.edu/home/pub/89/
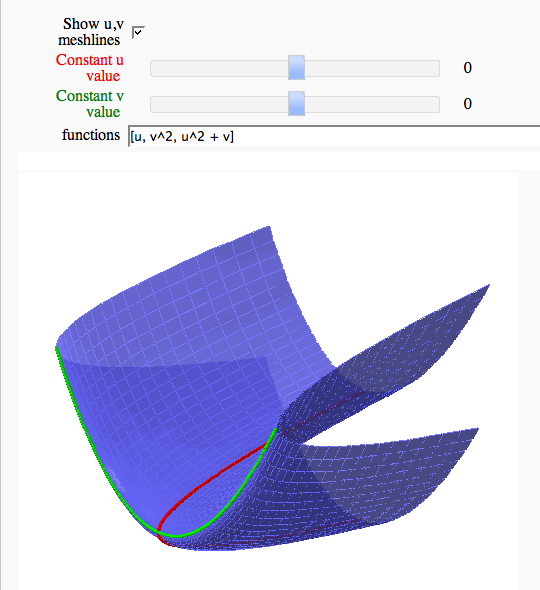
Parametric surface example (FIXME in Jupyter)
by Marshall Hampton
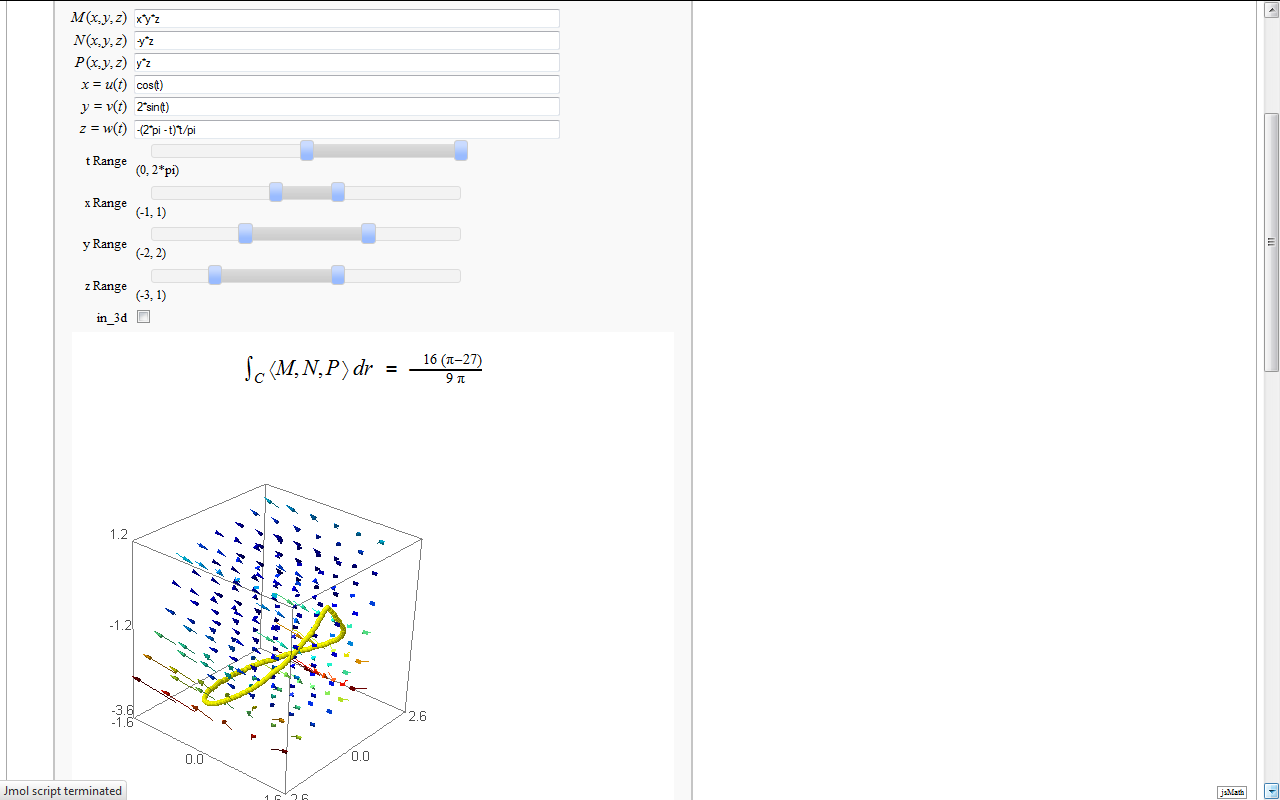
Line Integrals in 3D Vector Field
by John Travis