|
Size: 2819
Comment:
|
← Revision 11 as of 2020-02-08 13:26:23 ⇥
Size: 2898
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 6: | Line 6: |
| == Web app: protein browser == | == Web app: protein browser FIXME == |
| Line 8: | Line 8: |
| {{{ import urllib2 as U |
[sagecell-issues] {{{#!sagecell from six.moves.urllib.request import urlopen |
| Line 16: | Line 17: |
| f = U.urlopen(gen_str + GenBank_ID) g = f.read() f.close() |
with urlopen(gen_str + GenBank_ID) as f: g = f.read() |
| Line 25: | Line 25: |
| {{{ | [sagecell-issues] {{{#!sagecell |
| Line 57: | Line 58: |
| print 'Population size: ' + str(pop_size) print 'Selection coefficient for first taxon: ' + str(d_field(selection)) |
print('Population size: ' + str(pop_size)) print('Selection coefficient for first taxon: ' + str(d_field(selection))) |
| Line 68: | Line 69: |
| print 'Generations until coalescence: ' + str(len(gens)) show(coal_plot(coal_data1), axes = False, figsize = [8,4.0*len(gens)/pop_size], ymax = len(gens)-1) |
print('Generations until coalescence: ' + str(len(gens))) show(coal_plot(coal_data1), axes = False, figsize = [8, 4.0*len(gens)/pop_size], ymax = len(gens)-1) |
Sage Interactions - Bioinformatics
goto interact main page
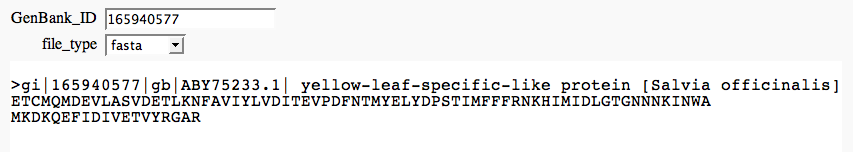
Web app: protein browser FIXME
by Marshall Hampton (tested by William Stein) [sagecell-issues]
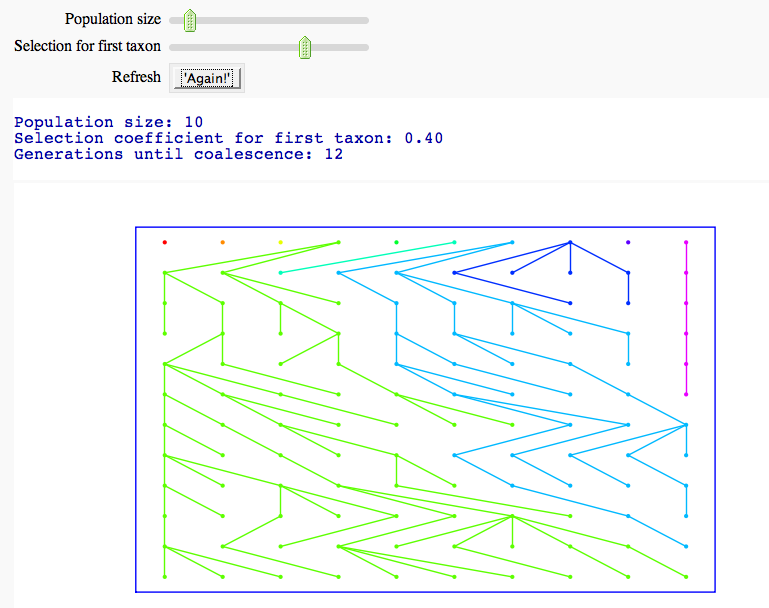
Coalescent simulator
by Marshall Hampton [sagecell-issues]